2024-08-20 ワシントン州立大学(WSU)
<関連情報>
- https://news.wsu.edu/press-release/2024/08/20/scientists-discover-new-code-governing-gene-activity/
- https://www.nature.com/articles/s41586-024-07662-z
ヒト塩基配列特異的転写因子の位置依存的機能 Position-dependent function of human sequence-specific transcription factors
Sascha H. Duttke,Carlos Guzman,Max Chang,Nathaniel P. Delos Santos,Bayley R. McDonald,Jialei Xie,Aaron F. Carlin,Sven Heinz & Christopher Benner
Nature Published:17 July 2024
DOI:https://doi.org/10.1038/s41586-024-07662-z
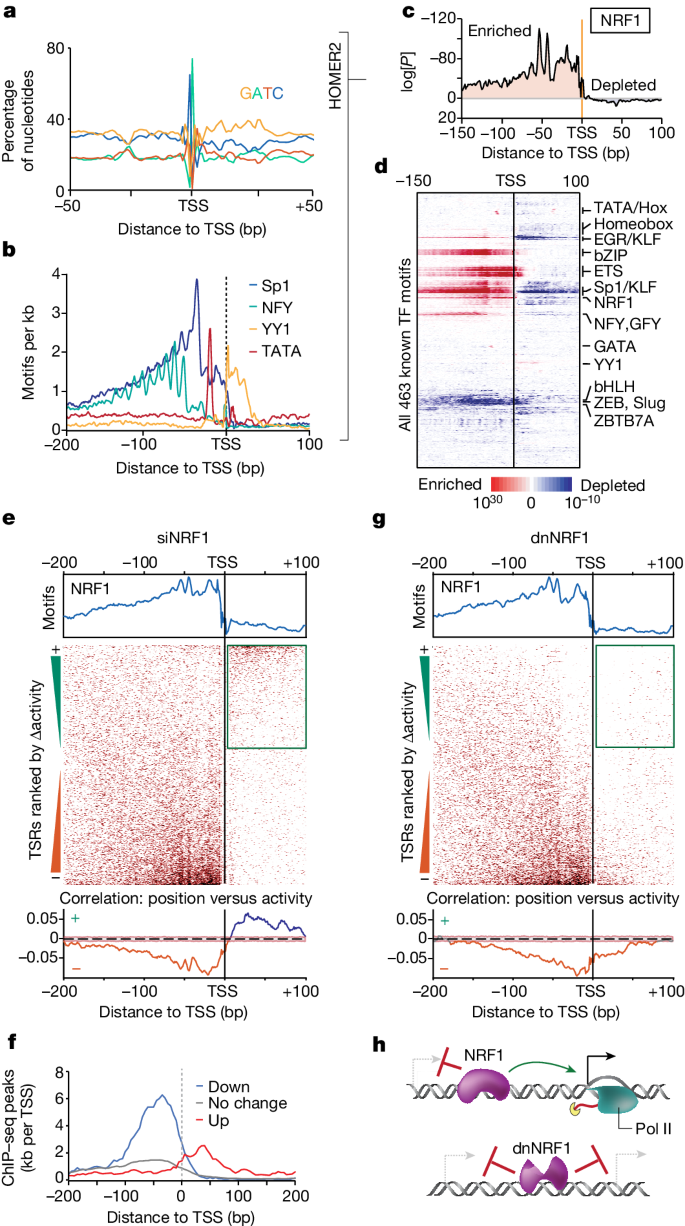
Abstract
Patterns of transcriptional activity are encoded in our genome through regulatory elements such as promoters or enhancers that, paradoxically, contain similar assortments of sequence-specific transcription factor (TF) binding sites1,2,3. Knowledge of how these sequence motifs encode multiple, often overlapping, gene expression programs is central to understanding gene regulation and how mutations in non-coding DNA manifest in disease4,5. Here, by studying gene regulation from the perspective of individual transcription start sites (TSSs), using natural genetic variation, perturbation of endogenous TF protein levels and massively parallel analysis of natural and synthetic regulatory elements, we show that the effect of TF binding on transcription initiation is position dependent. Analysing TF-binding-site occurrences relative to the TSS, we identified several motifs with highly preferential positioning. We show that these patterns are a combination of a TF’s distinct functional profiles—many TFs, including canonical activators such as NRF1, NFY and Sp1, activate or repress transcription initiation depending on their precise position relative to the TSS. As such, TFs and their spacing collectively guide the site and frequency of transcription initiation. More broadly, these findings reveal how similar assortments of TF binding sites can generate distinct gene regulatory outcomes depending on their spatial configuration and how DNA sequence polymorphisms may contribute to transcription variation and disease and underscore a critical role for TSS data in decoding the regulatory information of our genome.