2025-07-11 ミュンヘン大学(LMU)
<関連情報>
- https://www.lmu.de/en/newsroom/news-overview/news/ai-analysis-how-an-enzyme-associated-with-alzheimers-chooses-its-target-proteins.html
- https://www.nature.com/articles/s41467-025-60638-z
説明可能なAIによるγセクレターゼ基質のチャート化 Charting γ-secretase substrates by explainable AI
Stephan Breimann,Frits Kamp,Gabriele Basset,Claudia Abou-Ajram,Gökhan Güner,Kanta Yanagida,Masayasu Okochi,Stephan A. Müller,Stefan F. Lichtenthaler,Dieter Langosch,Dmitrij Frishman & Harald Steiner
Nature Communications Published:01 July 2025
DOI:https://doi.org/10.1038/s41467-025-60638-z
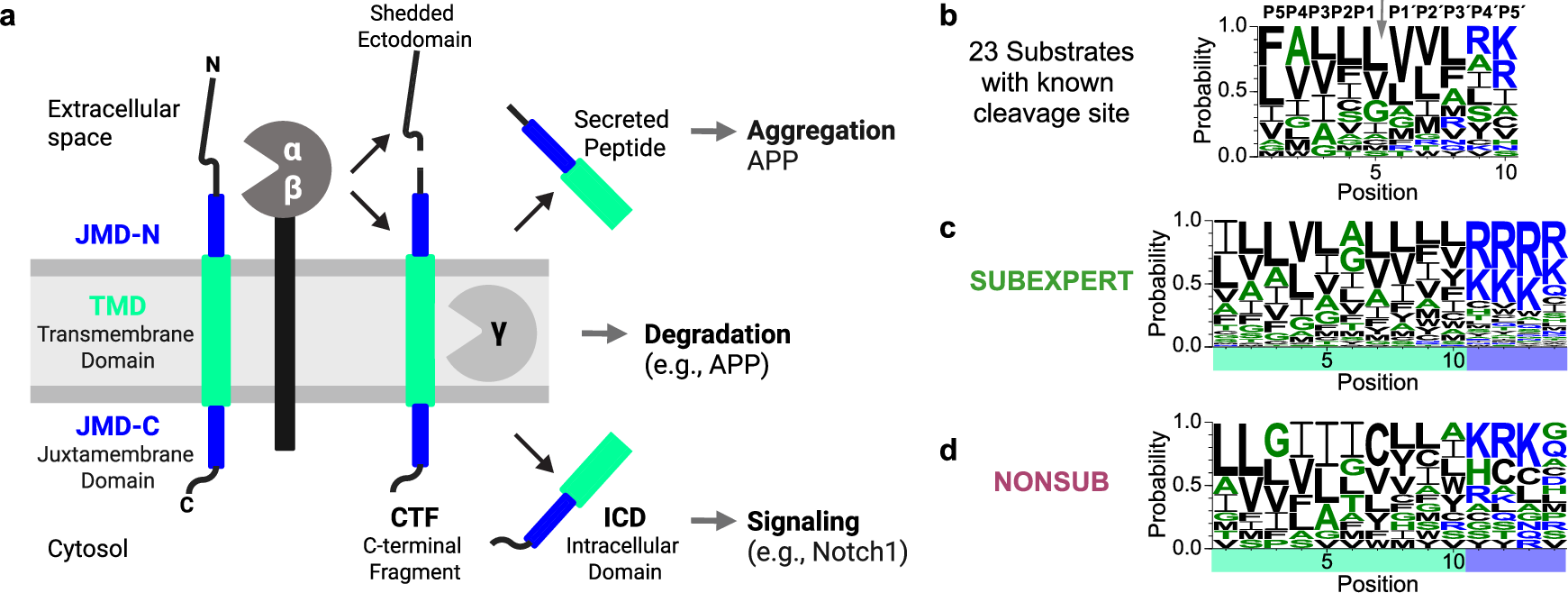
Abstract
Proteases recognize substrates by decoding sequence information—an essential cellular process elusive when recognition motifs are absent. Here, we unravel this problem for γ-secretase, an intramembrane-cleaving protease associated with Alzheimer’s disease and cancer, by developing Comparative Physicochemical Profiling (CPP), a sequence-based algorithm for identifying interpretable physicochemical features. We show that CPP deciphers a γ-secretase substrate signature with single-residue resolution, which can explain the conformational transitions observed in substrates upon γ-secretase binding. Using machine learning, we predict the entire human γ-secretase substrate scope, revealing numerous previously unknown substrates. Our approach outperforms state-of-the-art protein language models, improving prediction accuracy from 60% to 90%, and achieves an 88% success rate in experimental validation. Building on these advancements, we identify pathways and diseases not linked before to γ-secretase. Generally, CPP decodes physicochemical signatures—a concept that extends beyond sequence motifs. We anticipate that our approach will be broadly applicable to diverse molecular recognition processes.