2026-03-13 中国科学院(CAS)
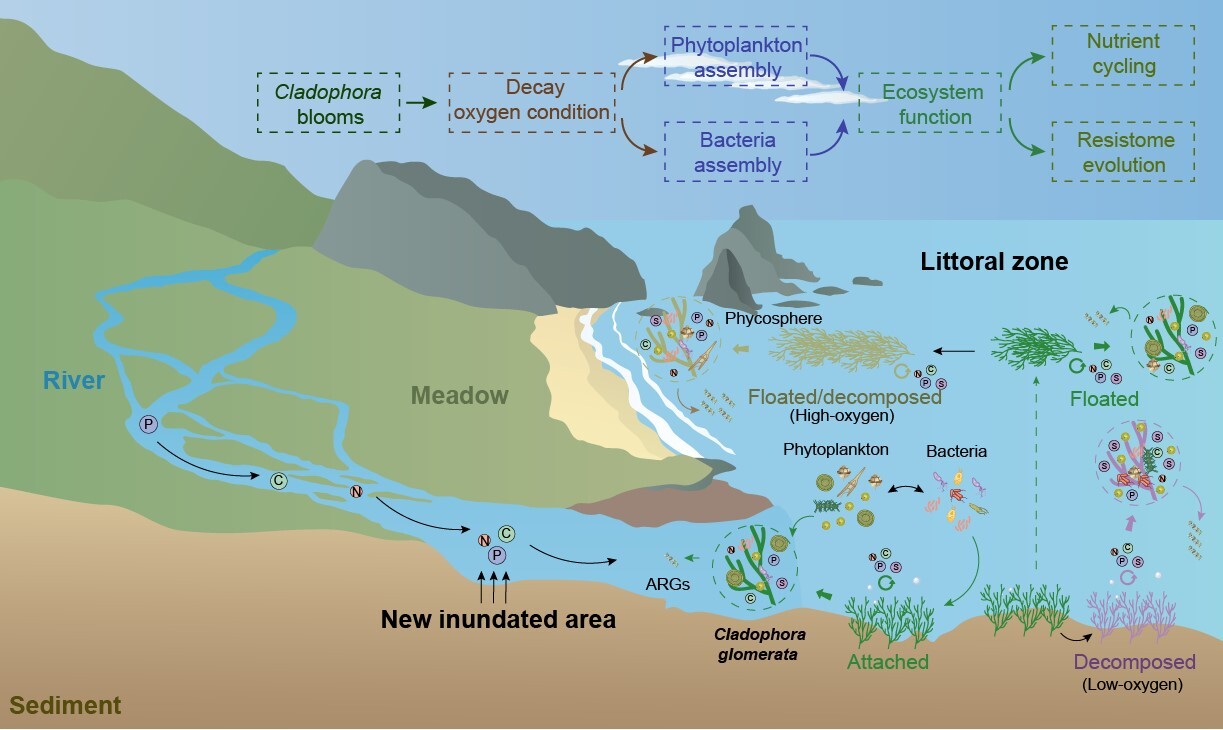
Effects of Cladophora blooms on nutrient cycling and antibiotic resistome evolution in the littoral zone of Qinghai Lake. (Image by IHB)
<関連情報>
- https://english.cas.cn/newsroom/research-news/202603/t20260313_1152667.shtml
- https://www.sciencedirect.com/science/article/abs/pii/S0043135426002125
青海湖沿岸域における着生植物群集と抗生物質耐性遺伝子群の進化は、クラドフォラ属によって促進される Cladophora drives the evolution of its epiphytic communities and antibiotic resistome in the littoral zone of Qinghai Lake
Jia Jia, Hongyi Ao, Xiong Xiong, Shuai Wang, Xiaoyan Xi, Kelong Chen, Chenxi Wu
Water Research Available online: 6 February 2026
DOI:https://doi.org/10.1016/j.watres.2026.125530
Highlights
- Cladophora blooms promote the proliferation of resistomes and nutrient cycling.
- The phycosphere acts as a hotspot for the transmission of resistance genes.
- Cladophora decay at low-oxygen conditions exhibits higher ecological risks.
- Resistome proliferation was primarily driven by VGT in the phycosphere.
- Cladophora-derived DOM shapes its epiphytic communities with a distance effect.
Abstract
Cladophora blooms, exacerbated by climate change and littoral eutrophication, pose a significant ecological threat. Of particular concern is their potential to disrupt phytoplankton and bacterial assemblages, triggering a cascade of effects that may include shifts in nutrient cycling and the dissemination of resistomes. However, the mechanistic links between Cladophora’s life-stage-dependent dissolved organic matter (DOM) release, its role in restructuring epiphytic communities, and its promotion of resistome dissemination in natural, oligotrophic lakes remain poorly understood. To address this, this study integrates field and laboratory investigations of Cladophora qinghaiensis sp. nov.. The algal phycosphere functions as a dynamic “gene incubator”, driven by chemical shifts in algal‑derived DOM. During decay under low‑oxygen conditions, DOM composition transitions from tyrosine‑like proteins to recalcitrant fulvic‑acid‑like compounds, selectively enriching competitive, intrinsically resistant taxa such as Halomonas and Phacus. Microbes such as Acinetobacter drive nutrient cycling (e.g., nitrogen metabolism) and serve as hotspots for resistomes within the phycosphere. Contrary to the expectation that high cell density favors horizontal gene transfer (HGT), genomic analyses show that vertical gene transfer (VGT) dominates antibiotic resistance gene (ARG) proliferation in this niche, a pattern explained by strong DOM‑mediated host selection and subsequent propagation. In contrast, the resistome in the surrounding water is more diverse and primarily shaped by HGT via mobile genetic elements. These results establish a mechanistic link between life‑stage‑specific algal DOM components, selective epiphytic communities enrichment, and divergent pathways of resistome evolution, positioning the phycosphere as a key source of ARGs that amplifies ecological risk in nearshore environments.


