2026-04-03 愛媛大学
<関連情報>
- https://www.ehime-u.ac.jp/data_relese/pr_20260403_pros/
- https://www.ehime-u.ac.jp/wp-content/uploads/2026/04/pr_20260331_pros.pdf
- https://www.cell.com/molecular-cell/fulltext/S1097-2765(26)00160-7
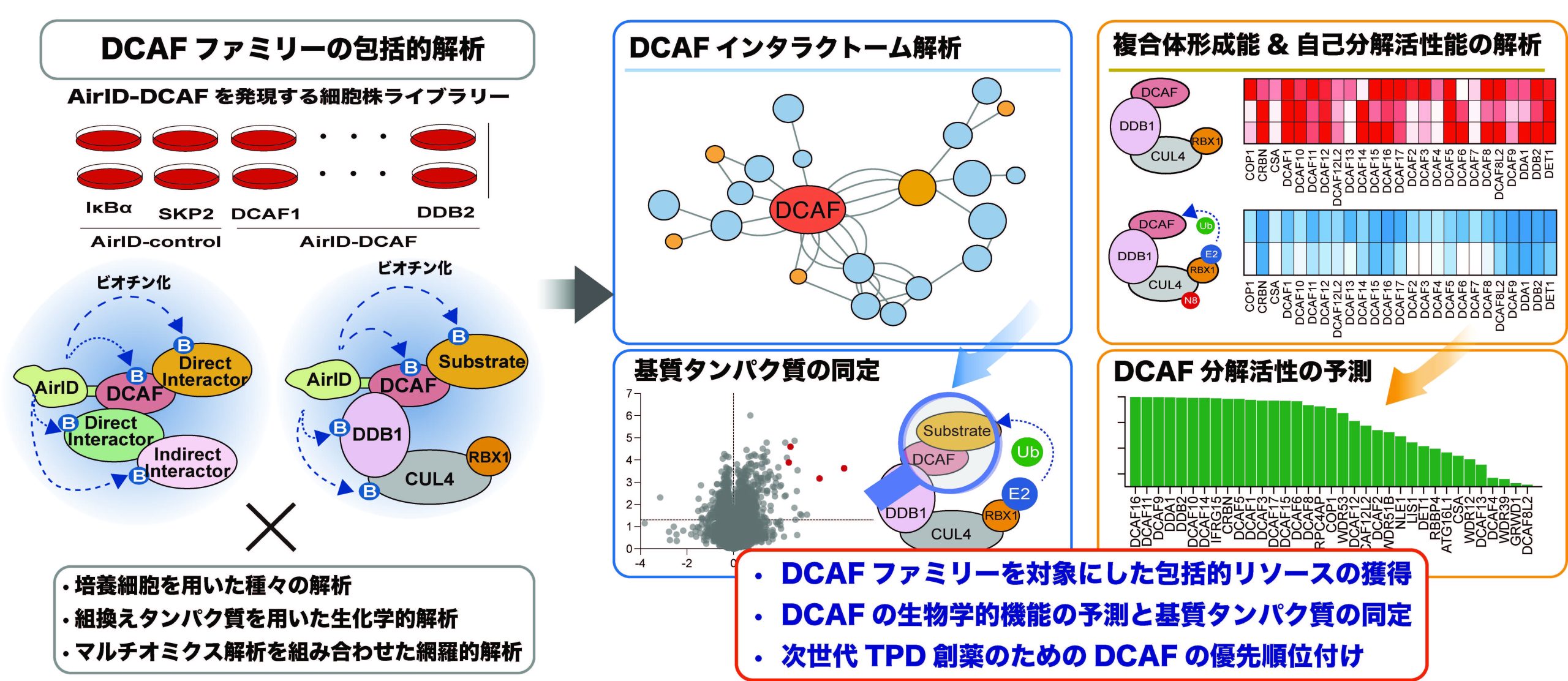
標的タンパク質分解におけるDDB1およびCUL4関連因子の優先順位付けのための相互作用ネットワークに基づくフレームワーク An interactome-based framework for DDB1- and CUL4-associated factor prioritization in targeted protein degradation
Satoshi Yamanaka ∙ Koya Nagaoka ∙ Yuki Shoya ∙ … ∙ Atsushi Hijikata ∙ Hidetaka Kosako ∙ Tatsuya Sawasaki
Molecular Cell Published: April 2, 2026
DOI:https://doi.org/10.1016/j.molcel.2026.03.004
Highlights
- A proximity interactome resource for the human DCAF family
- Systematic assessment of CRL4 formation and DCAF autodegradation
- Functional annotation and substrate inference from DCAF interactomes
- Prioritization of DCAFs for targeted protein degradation
Summary
The DDB1- and CUL4-associated factor (DCAF) family functions as substrate receptors within Cullin4-really interesting new gene (RING) ubiquitin ligases (CRL4s), facilitating proteasomal degradation of targeted substrates. Although CRL4-based targeted protein degradation (TPD) has emerged as a promising strategy to modulate undruggable proteins, the complex formation, substrates, and functional properties of many DCAFs remain poorly defined. In this study, using proximity biotinylation-based interactome analysis in human HEK293T cells, we systematically annotated interactors and functional associations of individual DCAFs. Furthermore, we identified substrates of the model DCAFs COP1 and DCAF3 using proximity biotinylation coupled with multi-omics approaches. By combining biochemical and cell-based analyses, we establish CRL4 complex formation and DCAF autodegradation as experimentally tractable proxy indicators to evaluate DCAF degradation activity and propose a set of high-activity DCAFs. These datasets establish a resource for functional characterization of DCAFs and provide a framework for their prioritization in TPD.