2025-02-25 ミュンヘン大学(LMU)
<関連情報>
- https://www.lmu.de/en/newsroom/news-overview/news/protein-design-flexible-components-allow-new-architectures.html
- https://www.nature.com/articles/s41594-025-01490-z
計算によって設計されたタンパク質集合体において、局所的な構造の柔軟性がオリゴモルフィズムを駆動する Local structural flexibility drives oligomorphism in computationally designed protein assemblies
Alena Khmelinskaia,Neville P. Bethel,Farzad Fatehi,Bhoomika Basu Mallik,Aleksandar Antanasijevic,Andrew J. Borst,Szu-Hsueh Lai,Ho Yeung Chim,Jing Yang ‘John’ Wang,Marcos C. Miranda,Andrew M. Watkins,Cassandra Ogohara,Shane Caldwell,Mengyu Wu,Albert J. R. Heck,David Veesler,Andrew B. Ward,David Baker,Reidun Twarock & Neil P. King
Nature Structural & Molecular Biology Published:26 February 2025
DOI:https://doi.org/10.1038/s41594-025-01490-z
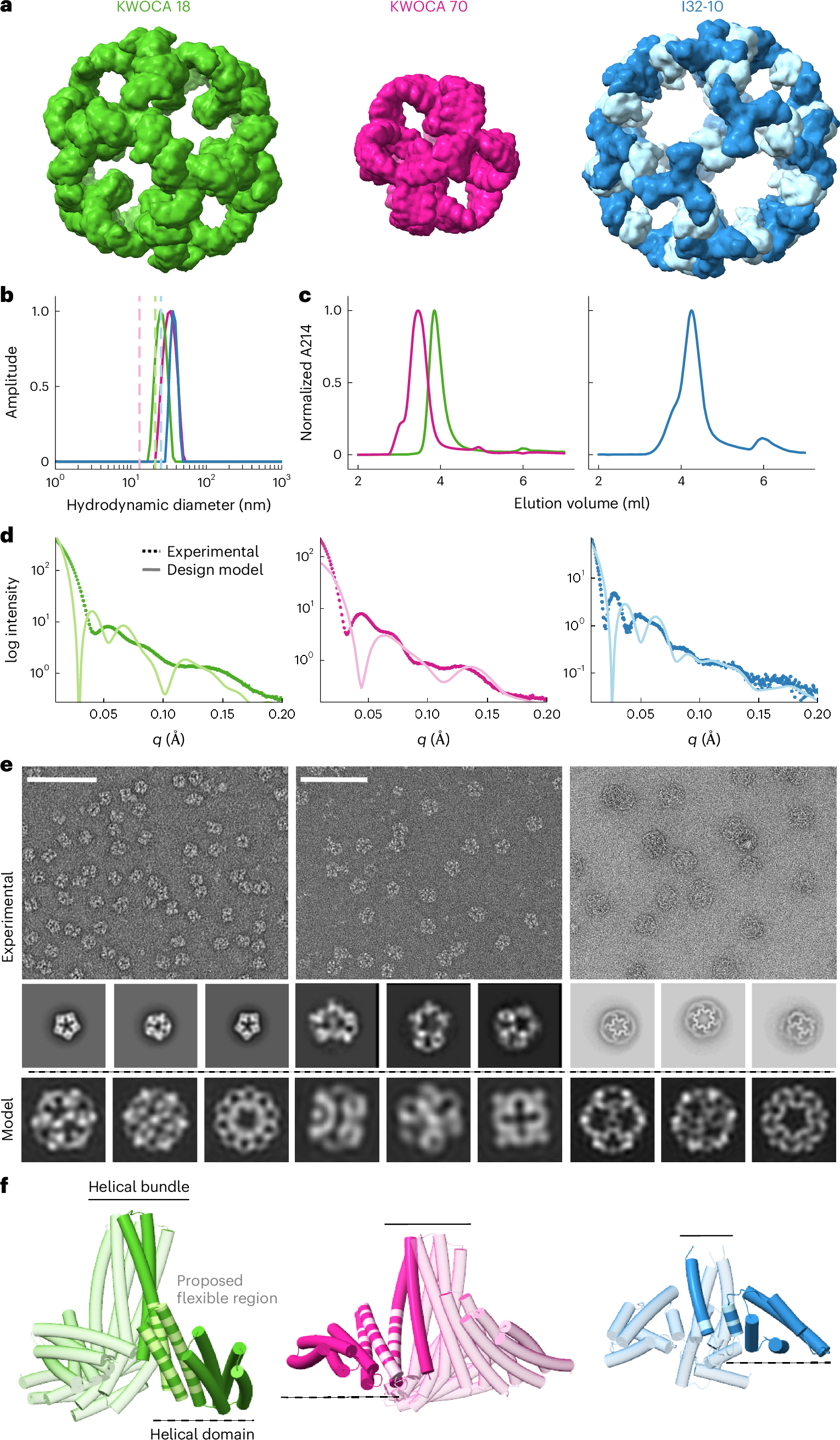
Abstract
Many naturally occurring protein assemblies have dynamic structures that allow them to perform specialized functions. Although computational methods for designing novel self-assembling proteins have advanced substantially over the past decade, they primarily focus on designing static structures. Here we characterize three distinct computationally designed protein assemblies that exhibit unanticipated structural diversity arising from flexibility in their subunits. Cryo-EM single-particle reconstructions and native mass spectrometry reveal two distinct architectures for two assemblies, while six cryo-EM reconstructions for the third likely represent a subset of its solution-phase structures. Structural modeling and molecular dynamics simulations indicate that constrained flexibility within the subunits of each assembly promotes a defined range of architectures rather than nonspecific aggregation. Redesigning the flexible region in one building block rescues the intended monomorphic assembly. These findings highlight structural flexibility as a powerful design principle, enabling exploration of new structural and functional spaces in protein assembly design.