2026-03-05 ロックフェラー大学
<関連情報>
- https://www.rockefeller.edu/news/39121-new-tracking-tool-reveals-how-t-cells-adapt-in-different-organs/
- https://www.nature.com/articles/s41590-026-02451-4
感染中のCD4⁺ T細胞の組織特異的クローン選択と分化 Tissue-specific clonal selection and differentiation of CD4⁺ T cells during infection
Roham Parsa,Helder C. Assis,Tiago B. R. de Castro,Gabriella Lima dos Reis,Arpita Sushil,Harald Hartweger,Angelina M. Bilate & Daniel Mucida
Nature Immunology Published:05 March 2026
DOI:https://doi.org/10.1038/s41590-026-02451-4
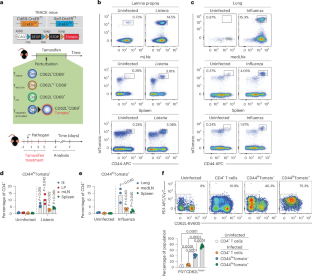
Abstract
Pathogen-specific CD4⁺ T cells expand and contract during infection, generating memory clones that shape subsequent immune responses. How distinct tissue environments influence differentiation and clonal selection of polyclonal T cells remains unclear. Here we develop Tracking Recently Activated Cell Kinetics (TRACK) mice, a dual-recombinase fate-mapping system enabling the spatial and temporal labeling of recently activated CD4⁺ T cells. Using TRACK mice during influenza infection, we observed organ-specific transcriptional differentiation and clonal selection in lung, mediastinal lymph nodes (medLNs) and spleen. During the effector phase, spleen-derived CD4⁺ T cells adopted a stem-like migratory phenotype, whereas medLN-activated cells differentiated into T follicular helper cells. T cell receptor sequencing showed low clonal overlap between tissues during the effector response, consistent with distinct antigenic landscapes. During memory formation, overlap increased between lung- and medLN-derived cells, while splenic clones retained a distinct repertoire. These findings define tissue-dependent mechanisms that shape CD4⁺ T cell fate and clonal architecture.


