2026-03-18 名古屋大学
<関連情報>
- https://www.nagoya-u.ac.jp/researchinfo/result/2026/03/post-964.html
- https://www.nagoya-u.ac.jp/researchinfo/result/upload_images/20260318_agr.pdf
- https://pubs.acs.org/doi/10.1021/acssynbio.5c00968
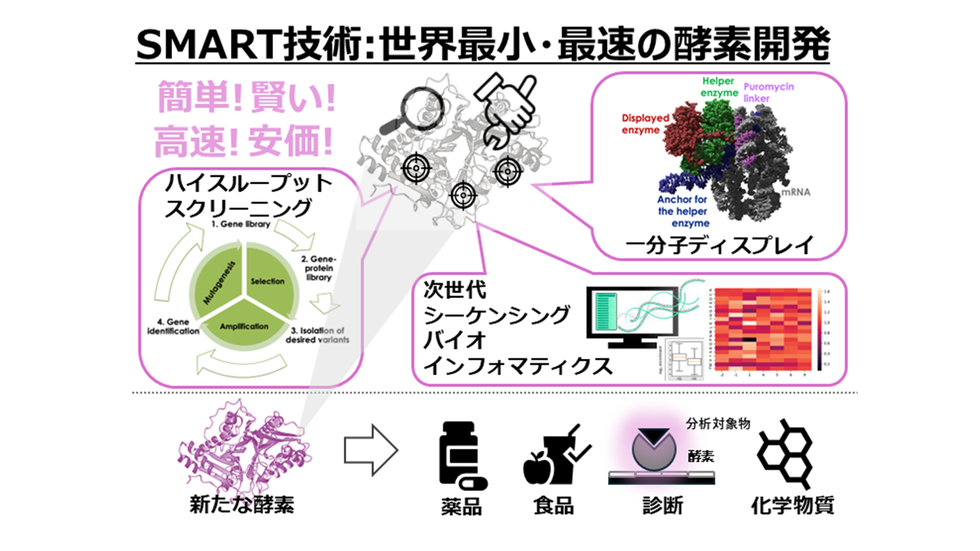
酵素進化のためのスマート単分子ディスプレイの力を活用する:オキシダーゼに焦点を当てて Harnessing the Power of SMART Single-Molecule Display for Enzyme Evolution: A Focus on Oxidase
Kalhari Munaweera,Nana Odake,Hannah Patricia Halim,Kakeru Ikeda,Bo Zhu,Maurizio Camagna,Tomokazu Ito,Tetsuya Kitaguchi,Naoto Nemoto,Hideo Nakano,and Jasmina Damnjanović
ACS Synthetic Biology Published: February 23, 2026
DOI:https://doi.org/10.1021/acssynbio.5c00968
Abstract
For a rapid and cost-effective evolution of tailor-made enzymes, we established a high-throughput in vitro selection platform named SMART (Single-Molecule Assay on Ribonucleic acid by Translated product), integrating mRNA display, next-generation sequencing, and bioinformatics. SMART represents a versatile system where a module termed an auxiliary unit allows enzyme-specific selection under various experimental conditions. Here, we report on the establishment of SMART for oxidases using a model enzyme, Schizosaccharomyces pombe d-amino acid oxidase (SpDAAO), and ascorbate peroxidase 2 as the auxiliary enzyme to detect hydrogen peroxide produced by the oxidase, and mediate biotinylation of active single-molecule display complexes. As a proof-of-concept, a library including site-saturation mutagenesis at the catalytic residue Y232 of SpDAAO was subjected to a single SMART selection round, yielding enrichment of the active enzyme variant. The results demonstrate the utility of SMART as a fast, robust, and efficient platform with the potential of customization for other enzyme chemistries through appropriate modifications of the auxiliary unit. Using SMART, desired enzyme variants can be selected in just a few hours by a single person without the need for costly equipment or any bias or limitations.