2026-03-19 マックス・プランク研究所
<関連情報>
- https://www.mpg.de/26286755/study-reveals-2fa-system-that-controls-protein-and-microrna-destruction
- https://www.nature.com/articles/s41586-026-10232-0
標的マイクロRNA分解を規定するE3ユビキチンリガーゼ機構 The E3 ubiquitin ligase mechanism specifying targeted microRNA degradation
Jakob Farnung,Elena Slobodyanyuk,Peter Y. Wang,Lianne W. Blodgett,Daniel H. Lin,Susanne von Gronau,Brenda A. Schulman & David P. Bartel
Nature Published:18 March 2026
DOI:https://doi.org/10.1038/s41586-026-10232-0
Abstract
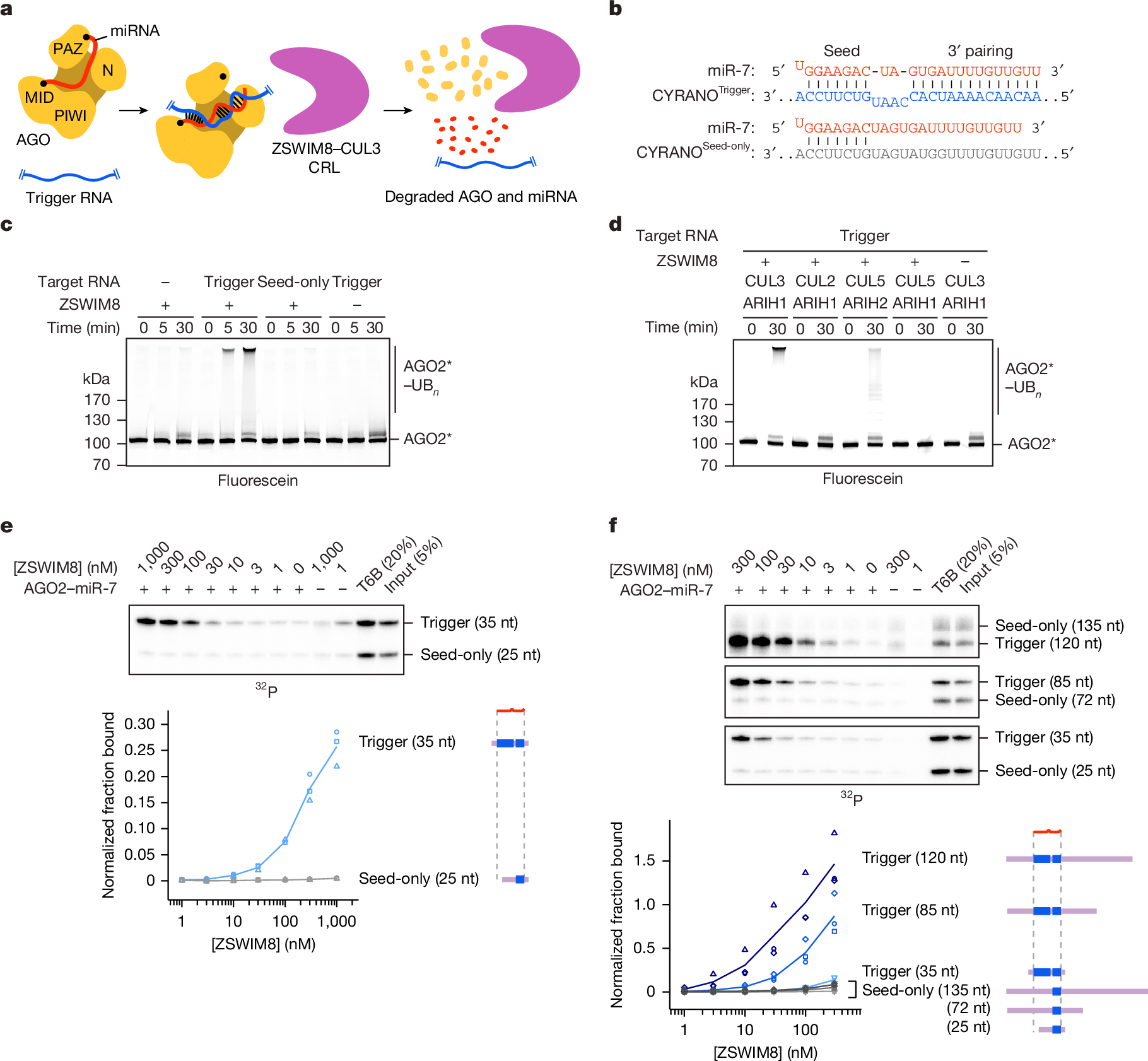
MicroRNAs (miRNAs) associate with Argonaute (AGO) proteins to form complexes that down-regulate target RNAs, including messenger RNAs from most human genes1,2,3. Within each complex, the miRNA pairs to target RNAs, and AGO provides effector function while also protecting the miRNA from cellular nucleases2,3,4,5. Although much is known about miRNA-directed gene regulation, less is known about how miRNAs themselves are regulated. One pathway that regulates miRNAs involves unusual targets called ‘trigger’ RNAs, which reverse the canonical regulatory logic and instead down-regulate miRNAs6,7,8,9. This target-directed miRNA degradation (TDMD) is thought to require a cullin–RING E3 ligase because it depends on the cullin protein CUL3 and other ubiquitylation components, including the BC-box protein ZSWIM8 (refs. 10,11). ZSWIM8 is required for murine perinatal viability and for destabilization of most short-lived miRNAs, which suggests biological importance of TDMD11,12,13. Here, biochemical and cellular assays establish AGO binding and polyubiquitylation by the ZSWIM8–CUL3 E3 ligase as the key regulatory steps of TDMD, and thereby define a unique cullin–RING E3 ligase class. Cryogenic electron microscopy analyses show ZSWIM8 recognizing distinct AGO and RNA conformations shaped by pairing of the miRNA to the trigger. Specificity of AGO ubiquitylation is established through generalizable RNA–RNA, RNA–protein and protein–protein interactions. The substrate features recognized by the E3 ligase do not conform to a conventional degron14,15 but instead establish a two-RNA-factor authentication mechanism for specifying a protein ubiquitylation substrate.

