2026-03-03 イリノイ大学アーバナ・シャンペーン校
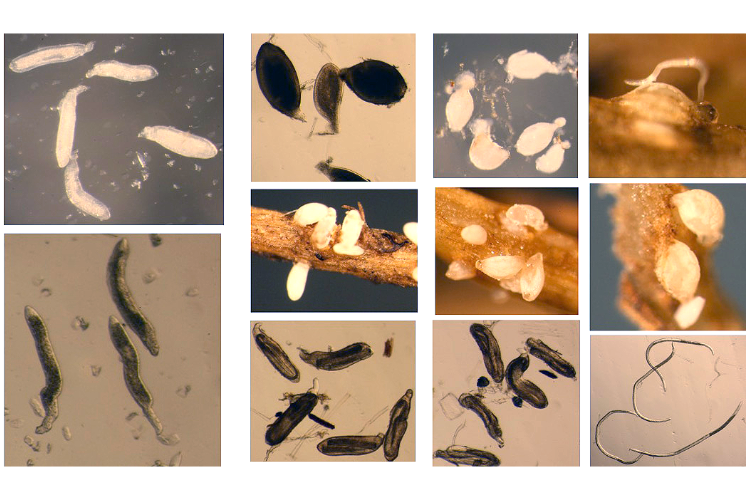
Soybean cyst nematode. Image credit Esmaeil Miraeiz.
<関連情報>
- https://aces.illinois.edu/news/comprehensive-genetic-library-soybean-cyst-nematode-could-renew-resistance-profitability
- https://link.springer.com/article/10.1186/s12864-025-12493-x
9つのダイズシストセンチュウゲノムのパンゲノム解析により、多様性と適応に寄与する隠れた変異が明らかになった Pangenome analysis of nine soybean cyst nematode genomes reveals hidden variation contributing to diversity and adaptation
Lucas Borges dos Santos,Kurt C. Showmaker,Rick E. Masonbrink,Kimberly K.O. Walden,João P. Gomes Viana,Khee-Man Kwon,Alvaro G. Hernandez,Zhihai Zhang,Christopher J. Fields,Thomas R. Maier,Andrew J. Severin,Thomas J. Baum,Melissa G. Mitchum & Matthew Hudson
BMC Genomics Published:15 January 2026
DOI:https://doi.org/10.1186/s12864-025-12493-x
Abstract
Background
The soybean cyst nematode (SCN) is a persistent threat to soybean production. SCN populations continually overcome resistant cultivars, causing significant yield losses. Studies conducted with a single reference genome restrict our understanding of intraspecific diversity, masking significant mechanisms of virulence evolution and host adaptation. Here we report a pangenome constructed of nine SCN populations of different pathotypes, including eight newly generated high-fidelity genome assemblies.
Results
We detected over 19,000 orthologous gene families and more than 12,000 putative secreted proteins in SCN. Combined, these data indicate substantial diversity across populations. Gene content analysis showed that 35% of gene families were the conserved core, 15% were soft-core, and 48% were accessory. Evidence of rapid evolution was identified in a high portion (40%) of core single-copy genes, most notably inside the protein domains responsible for host recognition and immune modulation. Analysis of gene-family expansion revealed extensive duplication and loss across lineages, suggesting ongoing paralog turnover within SCN populations. Finally, a graph-based pangenome enabled the identification of numerous structural variants within regions under selection.
Conclusions
Our study highlights substantial genetic variation in SCN that is not captured by single-reference analyses. By integrating multiple high-quality assemblies, we show that the SCN genome is highly dynamic, with extensive gene duplication and loss as well as structural variation shaping the differences among nematode populations. Collectively, the SCN pangenome provides a robust resource for studying virulence and adaptation mechanisms in SCN and establishes a genomic foundation for the development of more precise management strategies.


