2026-03-24 京都大学 iPS研究所
<関連情報>
- https://www.cira.kyoto-u.ac.jp/j/pressrelease/news/260324-100000.html
- https://www.nature.com/articles/s42003-026-09771-z
マイクロホモロジーを介した末端結合優性ターゲティングによるマウス胚におけるCRISPR精度の最適化 Optimizing CRISPR precision in mouse embryos via microhomology-mediated end joining-dominant targeting
Khanui Lkhagvadorj,Eiichi Okamura,Taito Taki,Hayate Suzuki,Akihiro Kuno,Yasushi Itoh,Seiya Mizuno,Knut Woltjen & Masatsugu Ema
Communications Biology Published:23 March 2026
DOI:https://doi.org/10.1038/s42003-026-09771-z
Abstract
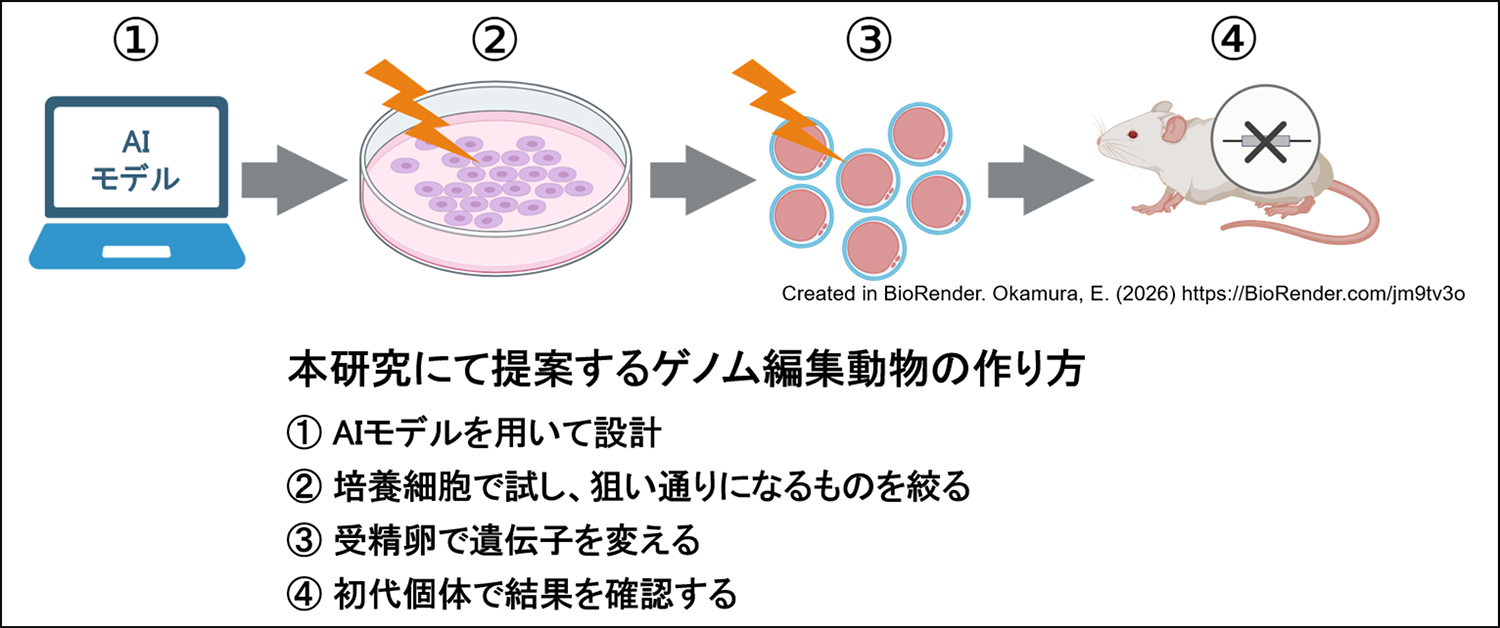
CRISPR/Cas9 technology enables efficient gene editing in mice, but its reliance on non-homologous end joining often leads to unpredictable and mosaic mutations in founder (F0) animals. Here, we present a hybrid genome editing strategy that combines in silico prediction software with in vitro validation using mouse embryonic stem cells (mESCs). Although the software was trained on mESC datasets, actual editing outcomes in mESCs more accurately reflected mutation patterns observed in blastocysts and post-implantation embryos. Using this information to develop an integrated pipeline, we pre-selected guide RNAs (gRNAs) predicted to promote microhomology-mediated end joining (MMEJ)-dominant repair and validated them in mESCs prior to embryo injection. Applied to the Tyr and Fgf10 genes, this approach enabled efficient generation of F0 mice with highly uniform genotypes. Our strategy enhances the predictability and reproducibility of CRISPR-based genome editing in mice and may help reduce animal usage in gene editing studies.