2022-05-06 スウェーデン・リンショーピング大学
<関連情報>
- https://liu.se/en/news-item/digital-twins-an-aid-to-give-individual-patients-the-right-treatment-at-the-right-time
- https://genomemedicine.biomedcentral.com/articles/10.1186/s13073-022-01048-4
デジタルツインを用いた動的な単一細胞による疾患遺伝子と創薬ターゲットの優先順位付けのフレームワーク A dynamic single cell-based framework for digital twins to prioritize disease genes and drug targets
Xinxiu Li,Eun Jung Lee,Sandra Lilja,Joseph Loscalzo,Samuel Schäfer,Martin Smelik,Maria Regina Strobl,Oleg Sysoev,Hui Wang,Huan Zhang,Yelin Zhao,Danuta R. Gawel,Barbara Bohle & Mikael Benson
Genome Medicine Published: 06 May 2022
DOI:https://doi.org/10.1186/s13073-022-01048-4
Abstract
Background
Medical digital twins are computational disease models for drug discovery and treatment. Unresolved problems include how to organize and prioritize between disease-associated changes in digital twins, on cellulome- and genome-wide scales. We present a dynamic framework that can be used to model such changes and thereby prioritize upstream regulators (URs) for biomarker- and drug discovery.
Methods
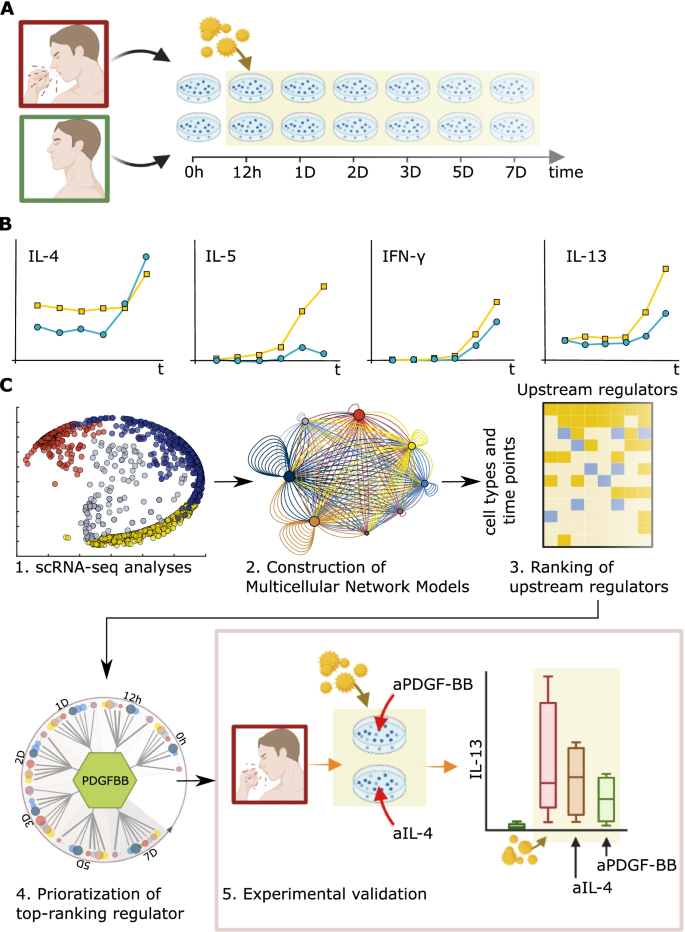
We started with seasonal allergic rhinitis (SAR) as a disease model, by analyses of in vitro allergen-stimulated peripheral blood mononuclear cells (PBMC) from SAR patients. Time-series a single-cell RNA-sequencing (scRNA-seq) data of these cells were used to construct multicellular network models (MNMs) at each time point of molecular interactions between cell types. We hypothesized that predicted molecular interactions between cell types in the MNMs could be traced to find an UR gene, at an early time point. We performed bioinformatic and functional studies of the MNMs to develop a scalable framework to prioritize UR genes. This framework was tested on a single-cell and bulk-profiling data from SAR and other inflammatory diseases.
Results
Our scRNA-seq-based time-series MNMs of SAR showed thousands of differentially expressed genes (DEGs) across multiple cell types, which varied between time points. Instead of a single-UR gene in each MNM, we found multiple URs dispersed across the cell types. Thus, at each time point, the MNMs formed multi-directional networks. The absence of linear hierarchies and time-dependent variations in MNMs complicated the prioritization of URs. For example, the expression and functions of Th2 cytokines, which are approved drug targets in allergies, varied across cell types, and time points. Our analyses of bulk- and single-cell data from other inflammatory diseases also revealed multi-directional networks that showed stage-dependent variations. We therefore developed a quantitative approach to prioritize URs: we ranked the URs based on their predicted effects on downstream target cells. Experimental and bioinformatic analyses supported that this kind of ranking is a tractable approach for prioritizing URs.
Conclusions
We present a scalable framework for modeling dynamic changes in digital twins, on cellulome- and genome-wide scales, to prioritize UR genes for biomarker and drug discovery.