2026-03-19 京都大学 iPS研究所
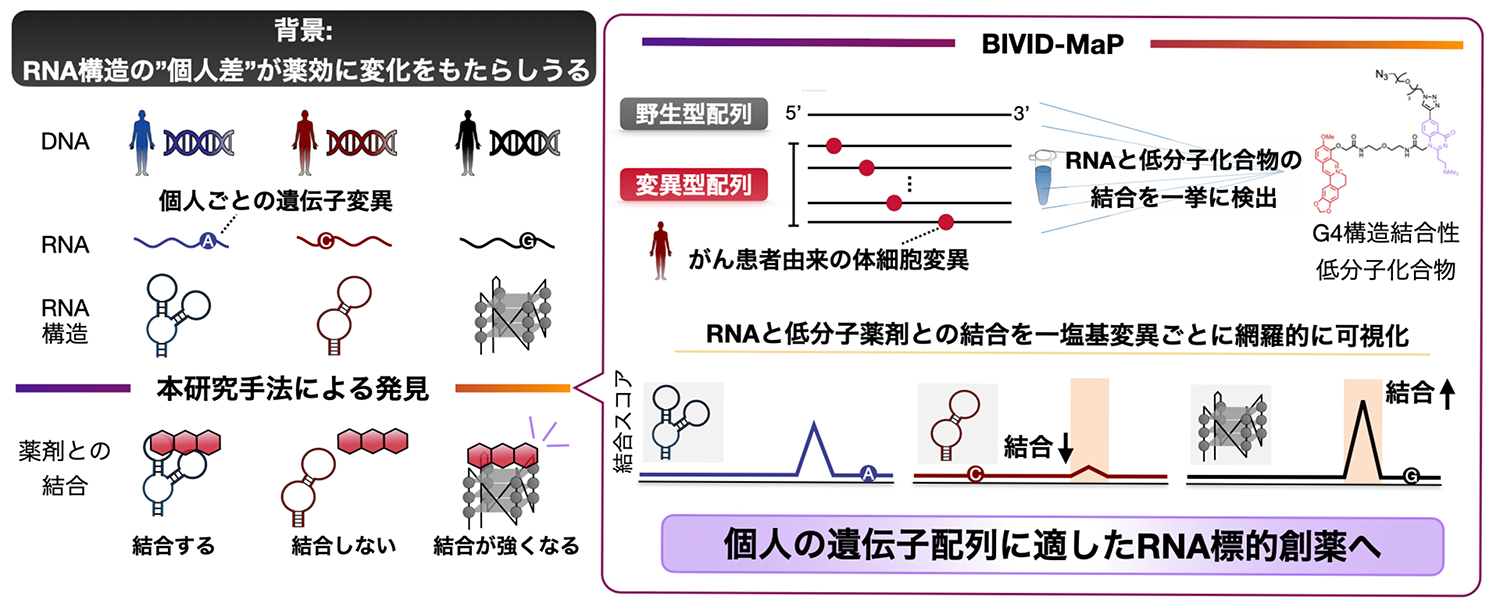
Fig. 1 本研究のまとめ
<関連情報>
- https://www.cira.kyoto-u.ac.jp/j/pressrelease/news/260319-190000.html
- https://www.nature.com/articles/s41467-026-70097-9
RNA G四重鎖を例とした、変異体特異的なRNA構造と低分子との相互作用の体系的な同定 Systematic identification of variant-specific RNA structure-small molecule interactions exemplified by RNA G-quadruplexes
Emi Miyashita,Kazumitsu Onizuka,Yutong Chen,Hiroki Yoshida,Hina Hatayama,Shunya Ishikawa,Peijie Yan,Takahito Hasegawa,Mamiko Ozawa,Kaho Maeta,Fumi Nagatsugi,Hirohide Saito & Kaoru R. Komatsu
Nature Communications Published:19 March 2026
DOI:https://doi.org/10.1038/s41467-026-70097-9
Abstract
Individual genetic variations, such as cancer-associated somatic mutations, alter RNA structures, thereby potentially enhancing or inhibiting the binding of RNA-targeting small molecules. However, to date, no approach has been available to identify these variant-specific RNA-small molecule interactions due to technical limitations. Here, we present Binding- and Vinyl-Quinazolinone-Induced Deletion-Based Mutational Profiling (BIVID-MaP), a high-throughput method for detecting RNA-small molecule interactions that combines binding-dependent covalent modification with profiling of deletions upon reverse transcription via deep sequencing. Using BIVID-MaP, we uncovered numerous variant-specific interactions between a G-quadruplex (G4)-binding small molecule and RNAs harboring single-nucleotide variants. Several cancer-associated somatic mutations significantly influence the binding intensity of a small molecule by affecting target G4 structures. These results demonstrate that BIVID-MaP can reveal previously ignored variant-specific RNA-small molecule interactions affected only by a single-nucleotide mutation, which may contribute to the development of RNA-targeting drugs in the future.