2026-03-02 ニューヨーク大学(NYU)
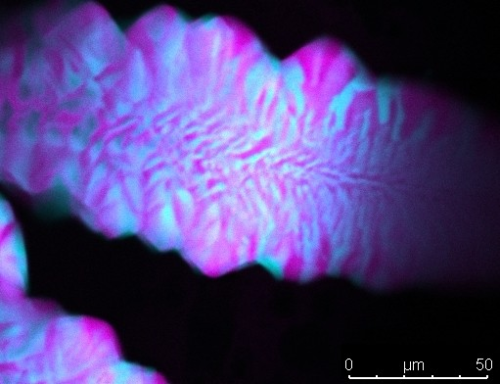
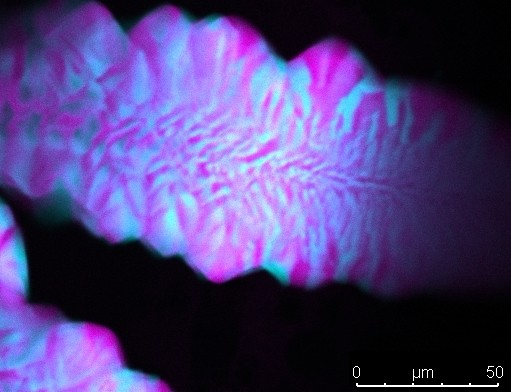
Right- and left-handed DNA tiles (blue and pink, respectively) are mixed to produce mirrored layers as a result of their shape and stacking. Image credit: Simon Vecchioni
<関連情報>
- https://www.nyu.edu/about/news-publications/news/2026/march/dna-structures.html
- https://www.nature.com/articles/s41467-026-69973-1
π-πスタッキングを用いたプログラム可能なDNA構造の鈍力アセンブリ Blunt-force assembly of programmable DNA architectures using π–π stacking
Karol Woloszyn,Andrew Horvath,Mara Jaffe,Lara Perren,Joe Rueb,Samyra Mahiba,Nataša Jonoska,Yoel P. Ohayon,James W. Canary,Simon Vecchioni & Ruojie Sha
Nature Communications Published:24 February 2026
DOI:https://doi.org/10.1038/s41467-026-69973-1 Unedited version
Abstract
Sticky-ended cohesion has been the driving force for DNA self-assembly and has enabled model and design programmability in DNA nanotechnology for over 40 years. Traditional units within self-assembled crystals have rationally-designed sticky ends to avoid unpredictable packing behavior, but in doing so, the surprising variety of contact flavors available to natural nucleic acids remains unexploited. Here, we employ composable DNA tiles to form complex 3D architectures using blunt-ended motifs with single duplex interfaces, thereby leveraging the geometry of the tile and the terminal nucleobase identity to control self-assembly outcomes. These crystals yielded X-ray diffraction at resolutions between 10.0 and 1.86 Å. We establish programmability and tunable packing, including translational and inversion symmetries, 5’−3’ and 5’−5’ stacking, and both positive and negative helical twist values. Finally, we establish the ability of racemic mixtures of L- and D-DNA motifs to perform programmable co-assembly driven by terminal π–π interactions, demonstrating molecular recognition and coexistence between mirror molecular systems.