2026-03-10 国立がん研究センター,東北大学
<関連情報>
- https://www.ncc.go.jp/jp/information/pr_release/2026/0310/index.html
- https://ashpublications.org/blood/article-abstract/doi/10.1182/blood.2025031466/566988/BCL11B-enhancer-hijacking-by-t-14-16-q32-q24
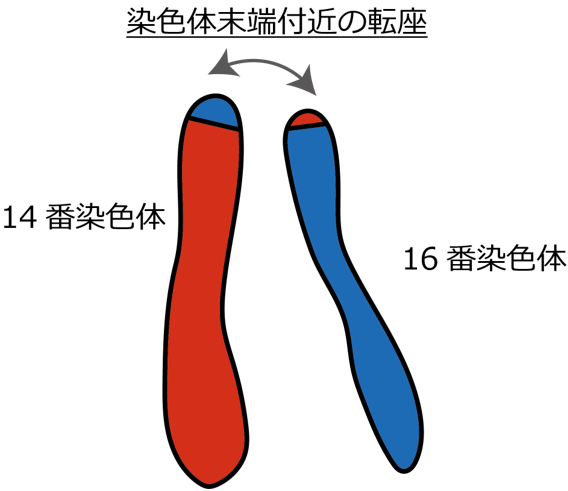
t(14;16)(q32;q24)転座によるBCL11Bエンハンサーハイジャックは、T-ALLの新たな高リスクサブタイプを定義する BCL11B enhancer hijacking by t(14;16)(q32;q24) translocation defines a novel high-risk subtype of T-ALL
Yoji Sasahara,Yoko Mizoguchi,Maiko Shimomura,Ryosuke Koyamada,Rintaro Ono,Daisuke Hasegawa,Kazuki Mitani,Hirohito Kubota,Satoshi Yoshihara,Nobuhiro Hiramoto,Akihito Otsuki,Yasunobu Okamura,Fumiki Katsuoka,Kengo Kinoshita,Masataka Hasegawa,Marina Togo-Ohno,Hirona Maeda,Nobuyuki Kakiuchi,Mai Takeuchi,Aiko Sato-Otsubo,Shota Kato,Kentaro Watanabe,Kotoe Katayama,Seiya Imoto,Yuichi Shiraishi,Katsuyoshi Koh,Souichi Suenobu,Eiso Hiyama,Susumu Goyama,Atsuo Kikuchi,Seishi Ogawa,Motohiro Kato,Yasuhito Nannya,Junko Takita,Kenichi Yoshida
Blood Published:March 6, 2026
DOI:https://doi.org/10.1182/blood.2025031466
Key Points
- We identified a novel T-ALL subtype with poor outcomes, overexpressing FENDRR, FOXF1, and FOXC2 through BCL11B enhancer hijacking.
- This subtype is characterized by frequent GATA3 mutations, marked lineage ambiguity, and enrichment in adolescents and young adults.
The molecular classification of T-cell acute lymphoblastic leukemia (T-ALL) remains incomplete, limiting risk stratification and the development of targeted therapies. Enhancer hijacking is a critical oncogenic mechanism that deregulates proto-oncogenes by repositioning cis-regulatory regions via structural variants. Here, we performed an integrated analysis of pediatric and adult T-ALL and mixed phenotype acute leukemias (MPALs), using whole-genome and whole-transcriptome sequencing. This analysis identified a group of 14 patients with predominantly T-lineage neoplasms driven by a t(14;16)(q32;q24) translocation, harboring universal GATA3 mutations and CDKN2A/B deletions. Mechanistically, this translocation repositions the ThymoD locus downstream of BCL11B, causing monoallelic, ectopic overexpression of FENDRR and mesenchymal transcription factor genes FOXF1 and FOXC2, activating epithelial-mesenchymal transition (EMT) transcription signatures. Immunophenotypic and single-cell RNA-seq analyses revealed marked lineage ambiguity with myeloid and B-cell differentiation potentials specific to this subtype. Furthermore, functional analyses in CD34-positive cord blood cells demonstrated that FOXF1 overexpression promotes myeloid differentiation while suppressing T-cell differentiation, serving as a key factor for lineage specification. Clinically, this subtype was detected in 0.15-4.0% of T-ALL/MPAL cases depending on the cohort, showing a median age of 15 years and enrichment in adolescents and young adults (AYA). Importantly, patients with t(14;16)(q32;q24) have an extremely poor prognosis, showing a trend toward worse outcomes than high-risk groups such as KMT2A-rearranged early T-cell progenitor (ETP)-like, SPI1-rearranged, and LMO2 γδ-like T-ALLs. The unique molecular landscape and poor prognosis of patients with the t(14;16)(q32;q24) translocation underscore the need for the development of novel subtype-specific therapeutic approaches.


