2026-03-13 産業技術総合研究所
<関連情報>
- https://www.aist.go.jp/aist_j/press_release/pr2026/pr20260313_3/pr20260313_3.html
- https://www.sciencedirect.com/science/article/pii/S0026265X26007253
マトリックス支援レーザー脱離イオン化イオンモビリティ質量分析法を用いた大規模研究のための超高スループットメタボロミクス Ultrahigh-throughput metabolomics for large-scale studies using matrix-assisted laser desorption/ionization-ion mobility-mass spectrometry
Hiroaki Takeda, Daisuke Miura, Teppei Shimamura, Yoshinori Fujimura, Masatomo Takahashi, Mitsuru Shindo, Ryo Nakabayashi, Takashi Nirasawa, Takeshi Bamba
Microchemical Journal Available online: 4 March 2026
DOI:https://doi.org/10.1016/j.microc.2026.117523
Highlights
- MALDI-IM-MS detects over 2000 metabolites within seconds per sample
- Quantitative reproducibility achieved with CV values below 20%
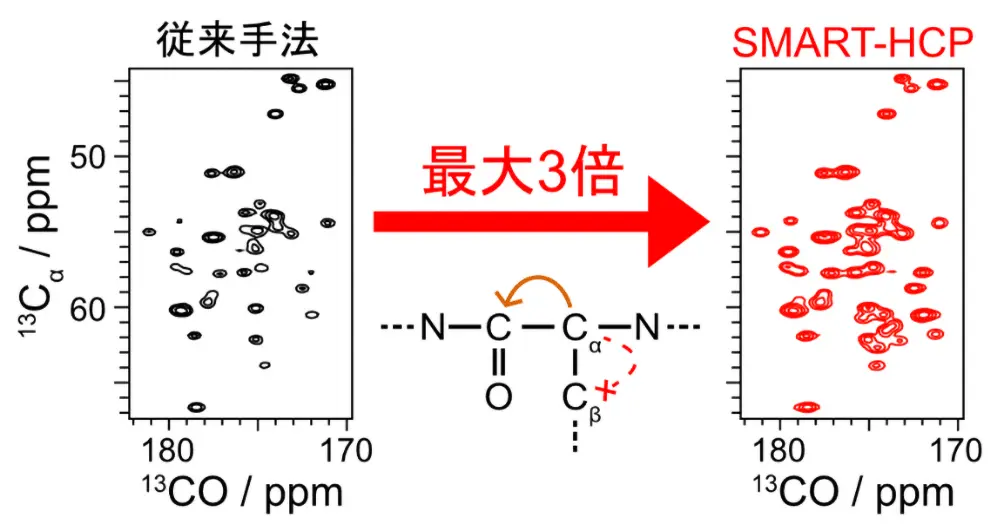
- Ion mobility with m/z enhances coverage and accuracy of detection
- Quantitative reproducibility achieved with CV values below 20%
- Plasma profiling visualizes metabolic differences among cancer types
Abstract
Background
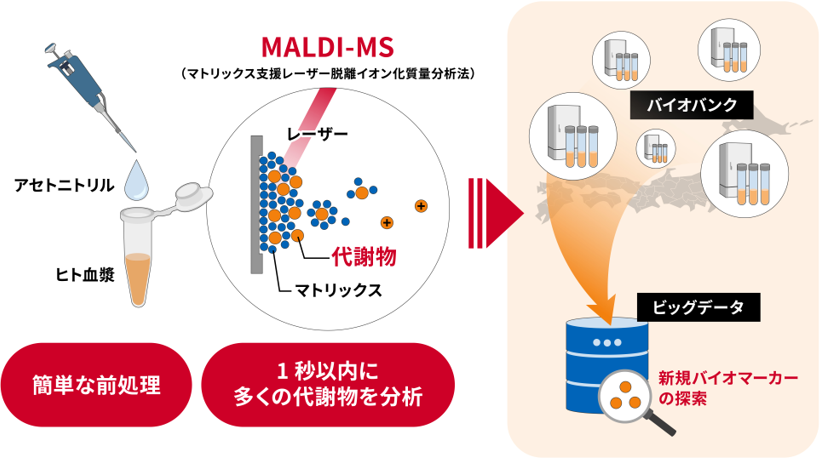
Metabolomics, the comprehensive analysis of low-molecular-weight metabolites, is central to biomarker discovery, drug development, and precision medicine. While large-scale metabolomics from biobanks could greatly advance individualized healthcare, conventional GC–MS and LC-MS approaches remain limited by time-consuming preparation and separation, restricting sample throughput. Matrix-assisted laser desorption/ionization mass spectrometry (MALDI-MS) offers a high-throughput alternative, being robust to contaminants and compatible with automated formats. Moreover, coupling MALDI with ion mobility (IM) expands molecular coverage by combining m/z and collision cross section measurements, positioning MALDI-IM-MS as a scalable platform for next-generation metabolomics and clinical big-data applications.
Results
The MALDI-IM-MS workflow, combined with acetonitrile extraction, enabled reproducible detection of >2000 features spanning hydrophilic and lipid metabolites across a broad m/z range. Ion mobility separation enhanced coverage beyond 700 features and resolved isomeric species by collision cross section (CCS) values. Averaging signals from multiple raster positions kept coefficients of variation below 20%, while adduct correlations (R2 > 0.90) indicated strong quantitative potential. UMAP analysis of 34 specimens reliably clustered replicates and separated healthy volunteers from cancer patients, revealing both shared and disease-specific metabolic alterations. Notably, plasma LPC levels, known to decrease in malignancies, were consistently reduced across patient groups, confirming clinical observations. Despite the relatively small cohort, these results demonstrate that MALDI-IM-MS can sensitively capture disease-related metabolic alterations and provides a scalable platform for large-scale cancer metabolomics and precision medicine applications.
Significance
The approach presented in this study offers a promising solution for conducting large-scale clinical trials with minimal effort, and is expected to serve as a big data generation platform that contributing to “precision medicine” by utilizing existing biobanks. Furthermore, we believe that this technology can be used not only in clinical research, but also to large-scale chemical and biochemical screening in a wide range of research fields, including pharmaceuticals, food, fisheries, livestock, and agriculture.