2026-04-13 京都大学
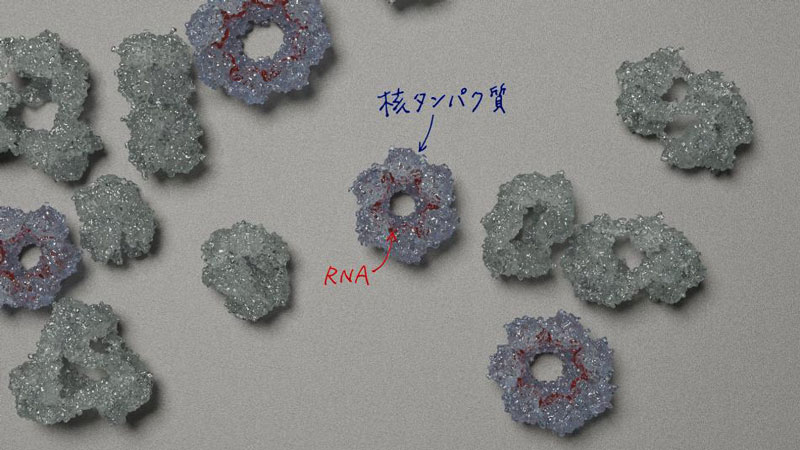

本研究の概念図:多数の複合体状態から、核タンパク質-RNA複合体の構造を見出した。©杉田征彦
<関連情報>
- https://www.kyoto-u.ac.jp/ja/research-news/2026-04-13
- https://www.kyoto-u.ac.jp/sites/default/files/2026-04/web_2604_Sugita-205b5213b5653d782fbe21afa7755328.pdf
- https://www.science.org/doi/10.1126/sciadv.aeb0835
ボルナ病ウイルス1型核タンパク質–RNA複合体の構造と形成過程 Structure and assembly of Borna disease virus 1 nucleoprotein-RNA complexes
Yukihiko Sugita, Yuya Hirai, Shinya H. Goto, Takuro Fujiwara, […] , and Masayuki Horie
Science Advances Published:10 Apr 2026
DOI:https://doi.org/10.1126/sciadv.aeb0835
Abstract
Structures of nucleoprotein (N)–RNA complexes of the Bornaviridae, a virus family in the order Mononegavirales, have not been reported. Here, using cryo–electron microscopy (cryo-EM), we report high-resolution structures of Borna disease virus 1 (BoDV-1) N-RNA complex assemblies, including a dominant hexameric ring-like complex and less populated heptameric and octameric forms, the first RNA-bound N structures reported from this family. These structures reveal key features of N-RNA engagement and a BoDV-1–specific stoichiometry of eight nucleotides per N, providing a framework for comparison with related negative-strand RNA viruses. In addition to these RNA-bound complexes, we identified multiple RNA-free oligomers, indicating substantial conformational flexibility of N. Mutational analyses identified residues essential for nucleocapsid formation and RNA synthesis. Cryo-EM of mutant complexes captured RNA-free assemblies, suggesting that initial N oligomerization precedes RNA binding. These findings clarify the structural organization of the N-RNA complex and suggest how oligomeric plasticity contributes to nucleocapsid assembly.