2026-04-15 ロックフェラー大学
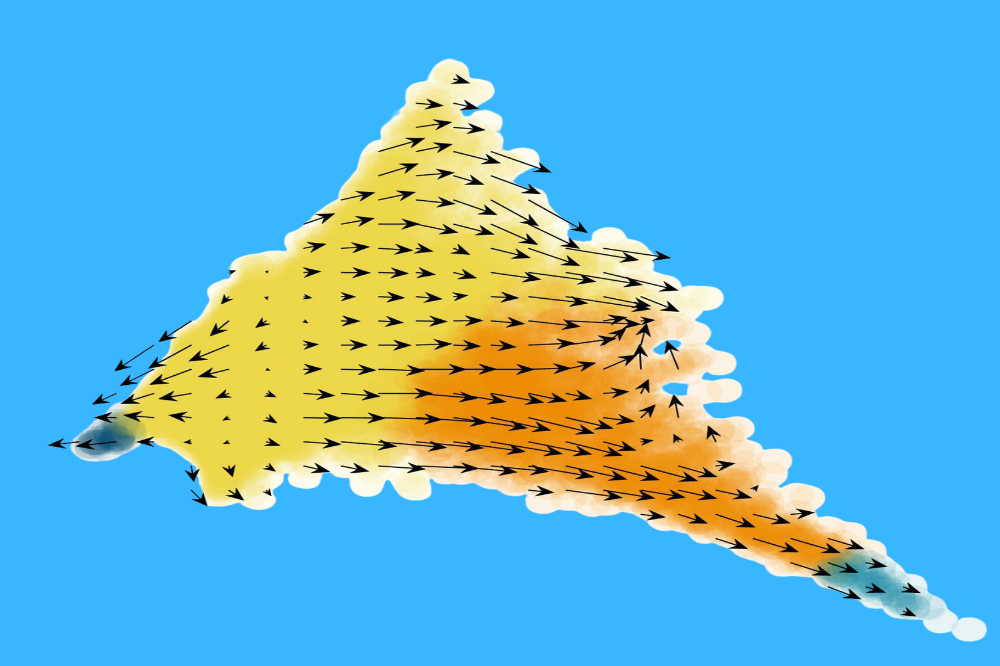

Cell state transition trajectories from genetically perturbed cells. (Credit: Cao lab)
<関連情報>
- https://www.rockefeller.edu/news/39376-novel-tool-could-identify-new-therapeutic-targets-in-complex-diseases-like-cancer/
- https://www.nature.com/articles/s41586-026-10367-0
PerturbFateによるメラノーマ薬剤耐性の収束制御因子のマッピング Mapping convergent regulators of melanoma drug resistance by PerturbFate
Zihan Xu,Ziyu Lu,Aileen Ugurbil,Abdulraouf Abdulraouf,Andrew Liao,Jianxiang Zhang,Wei Zhou & Junyue Cao
Nature Published:15 April 2026
DOI:https://doi.org/10.1038/s41586-026-10367-0
Abstract
High-throughput genomic studies have uncovered associations between diverse genetic alterations and disease phenotypes. However, elucidating how perturbations in functionally disparate genes give rise to convergent cellular states remains challenging. Here we present PerturbFate, a high-throughput, cost-effective, combinatorial-indexing single-cell platform that enables systematic interrogation of massively parallel CRISPR interference1 perturbations across the full spectrum of gene regulation, from chromatin remodelling and nascent transcription to steady-state transcriptomic phenotypes. Using PerturbFate, we profiled more than 300,000 cultured melanoma cells to characterize multimodal phenotypic and gene regulatory responses to perturbations in more than 140 vemurafenib resistance-associated genes. We uncovered a shared dedifferentiated cell state marked by convergent cooperative transcription factor activities across diverse genetic perturbations. We further dissected phenotypic responses to perturbations in Mediator complex components, linking module-specific biochemical properties to convergent transcriptional activations. We identified common regulatory nodes that drive similar phenotypic outcomes across distinct genetic perturbations. We also delineated how perturbations in functionally unrelated genes reshape cell state. Thus, PerturbFate establishes a versatile platform for identifying key molecular regulators by anchoring multimodal regulatory dynamics to disease-relevant phenotypes.