20026-04-22 バーミンガム大学
<関連情報>
- https://www.birmingham.ac.uk/news/2026/breastfed-babies-balance-bacteria-in-developing-microbiome
- https://www.nature.com/articles/s41467-026-71764-7
- https://academic.oup.com/ismej/article/14/2/635/7474845
ヒト母乳オリゴ糖は、大腸菌とビフィドバクテリウム・ビフィダム間の共生関係を媒介する Human milk oligosaccharide mediates mutualism between Escherichia coli and Bifidobacterium bifidum
David Seki,Shaul Pollak,Magdalena Kujawska,Raymond Kiu,Antia Acuna-Gonzalez,Lucy I. Crouch,Cassie R. Bakshani,Peter T. Chivers,Monique Mommers,Nils van Best,John Penders & Lindsay J. Hall
Nature Communications Published:22 April 2026
DOI:https://doi.org/10.1038/s41467-026-71764-7
Abstract
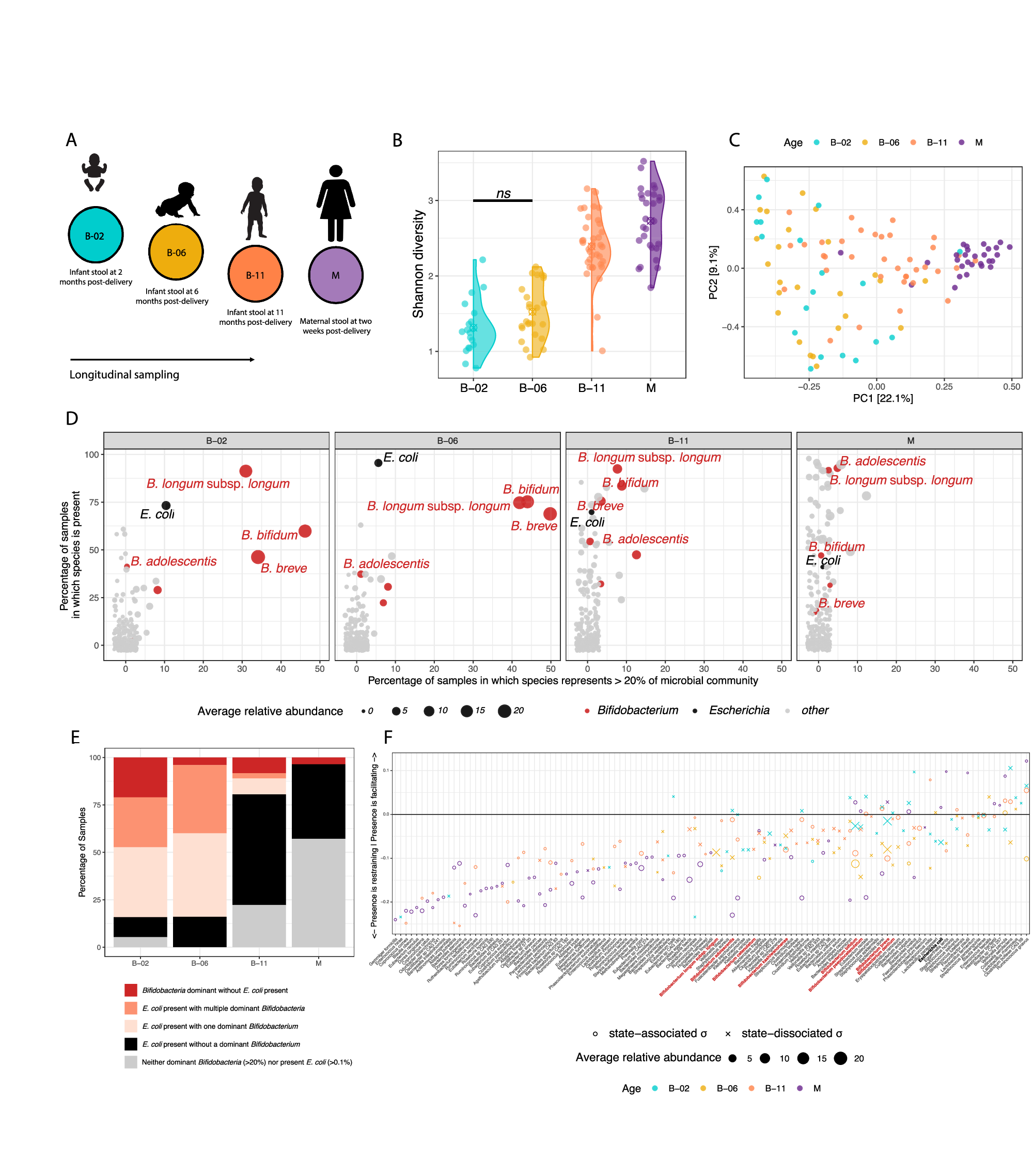
Infant gut microbiota development involves frequent colonization by Enterobacteriaceae, particularly Escherichia coli, yet their ecological role in healthy infants is unclear. Here, we analyse longitudinal stool samples from healthy, term-born, breastfed infants (n = 41) and related mothers (n = 30) using shotgun metagenomics and novel computational approaches. Strain-resolved profiling indicates that Bifidobacterium species are frequently shared within families, whereas E. coli derive from external sources, but often persist within individuals. Despite differing ecological strategies, these genera co-exist and share evolutionary adaptations related to lactose acquisition in the infant gut. In vitro, we demonstrate that interactions between E. coli and Bifidobacterium bifidum are mutualistic in co-culture, where E. coli supplies cysteine to its auxotrophic partner, facilitating cooperative degradation of 2′-fucosyllactose, the predominant human milk oligosaccharide. In turn, the liberated monosaccharides sustain E. coli growth, highlighting a cooperative cross-feeding interaction that may contribute to regulating E. coli abundance within the infant host.
母乳由来のヒトミルクオリゴ糖は、単一の生態系内での ビフィズス菌の相互作用を促進する Breast milk-derived human milk oligosaccharides promote Bifidobacterium interactions within a single ecosystem
Melissa A E Lawson,Ian J O’Neill,Magdalena Kujawska,Sree Gowrinadh Javvadi,Anisha Wijeyesekera,Zak Flegg,Lisa Chalklen,Lindsay J Hall
The ISME Journal Published:18 November 2019
DOI:https://doi.org/10.1038/s41396-019-0553-2
Abstract
Diet-microbe interactions play an important role in modulating the early-life microbiota, with Bifidobacterium strains and species dominating the gut of breast-fed infants. Here, we sought to explore how infant diet drives distinct bifidobacterial community composition and dynamics within individual infant ecosystems. Genomic characterisation of 19 strains isolated from breast-fed infants revealed a diverse genomic architecture enriched in carbohydrate metabolism genes, which was distinct to each strain, but collectively formed a pangenome across infants. Presence of gene clusters implicated in digestion of human milk oligosaccharides (HMOs) varied between species, with growth studies indicating that within single infants there were differences in the ability to utilise 2′FL and LNnT HMOs between strains. Cross-feeding experiments were performed with HMO degraders and non-HMO users (using spent or ‘conditioned’ media and direct co-culture). Further 1H-NMR analysis identified fucose, galactose, acetate, and N-acetylglucosamine as key by-products of HMO metabolism; as demonstrated by modest growth of non-HMO users on spend media from HMO metabolism. These experiments indicate how HMO metabolism permits the sharing of resources to maximise nutrient consumption from the diet and highlights the cooperative nature of bifidobacterial strains and their role as ‘foundation’ species in the infant ecosystem. The intra- and inter-infant bifidobacterial community behaviour may contribute to the diversity and dominance of Bifidobacterium in early life and suggests avenues for future development of new diet and microbiota-based therapies to promote infant health.


