2026-03-30 国立遺伝学研究所
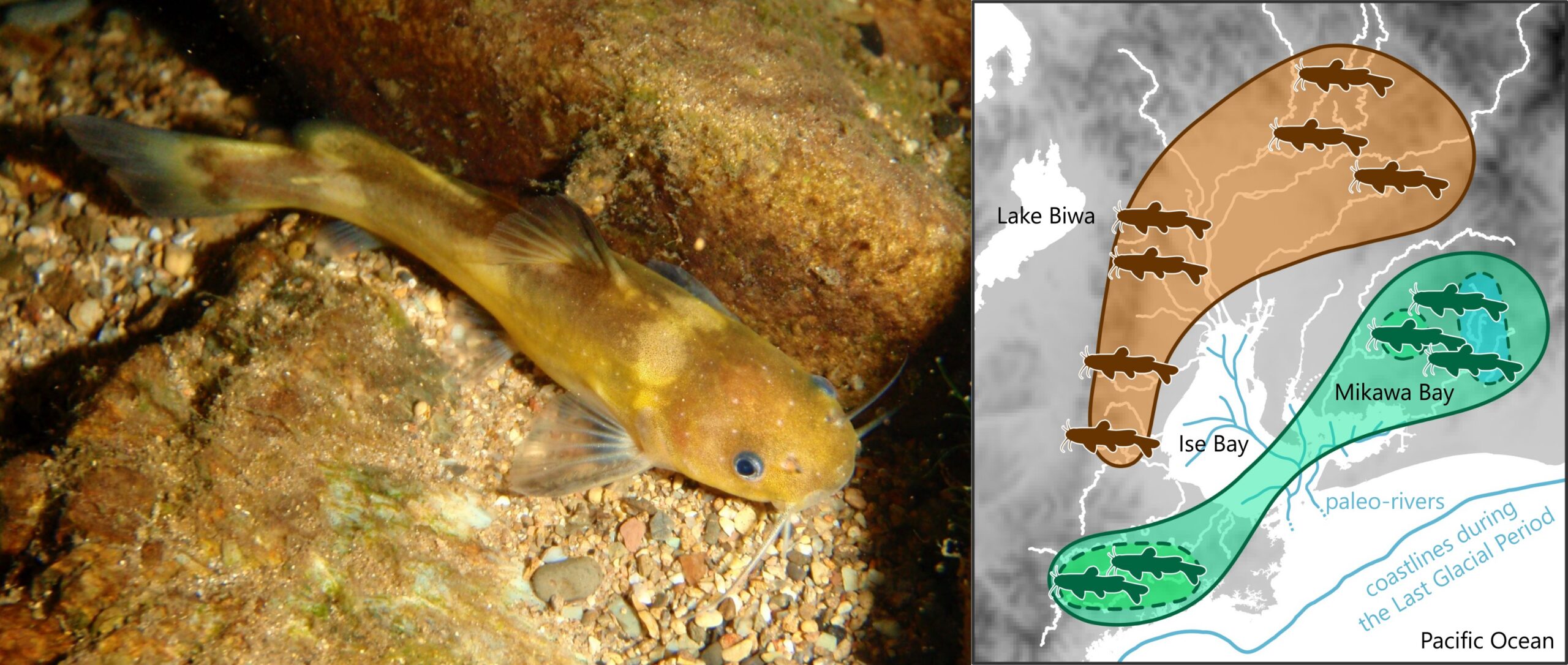

ネコギギ(撮影:渡辺勝敏教授)と本研究で推定された本種の地域集団間の遺伝的関係。白線は現在の河川,青線は最終氷期の古水系と海岸線をそれぞれ表す。
<関連情報>
- https://www.nig.ac.jp/nig/ja/2026/03/research-highlights_ja/pr20260330_2.html
- https://www.nig.ac.jp/nig/images/research_highlights/PR20260330_2.pdf
- https://onlinelibrary.wiley.com/doi/10.1002/ece3.73263
ゲノムワイド解析により、絶滅危惧種であるイチカワイナマズ(Pseudobagrus ichikawai)の最近の分岐によって形成された集団構造が解明された Genome-Wide Analysis Successfully Resolves Population Structure Shaped by Recent Divergence in the Endangered Bagrid Catfish Pseudobagrus ichikawai
Keisuke Onuki, Ryoichi Tabata, Tappei Mishina, Mutsumi Nishida, Katsutoshi Watanabe
Ecology and Evolution Published: 26 March 2026
DOI:https://doi.org/10.1002/ece3.73263
ABSTRACT
The population structure and history of the endangered bagrid catfish Pseudobagrus ichikawai were assessed to provide insights into its conservation through comparative analyses of several traditional genetic markers and genome-wide SNP data. The species is distributed in the Ise Bay area, located in central Honshu Island, Japan, a biogeographically unique region with endemic freshwater species. Samples were collected across all major populations, and genetic differentiation, population history, and demographic trends were inferred. Genetic analyses were conducted using whole mitochondrial genome sequences (mitogenomes), microsatellite polymorphisms, reduced-representation genome sequencing (MIG-seq), and whole-genome resequencing. For the latter two methods, the newly determined chromosome-level genome assembly was used as a reference. Genome-wide data revealed a pattern of population differentiation that could not be detected with mitogenome or microsatellite data. This differentiation pattern, including genetic similarity between populations across the eastern and western sides of Ise Bay (trans-Ise Bay pattern), appears to reflect population connectivity facilitated by paleo-river systems during the Last Glacial Period. Genetic diversity indices showed low variability across populations, suggesting historical bottlenecks and, for some populations, recent inbreeding. By utilizing genome-wide data, this study elucidated the subtle population structure of P. ichikawai in unprecedented detail. This approach provides a deeper understanding of the species’ population history under geographic and climatic influences, contributing to conservation strategies for regional biodiversity management. Our results also demonstrate that the absence of detectable population structure using traditional markers does not necessarily preclude its existence, highlighting the critical role of genome-wide data in uncovering cryptic diversity.