2026-03-31 沖縄科学技術大学院大学
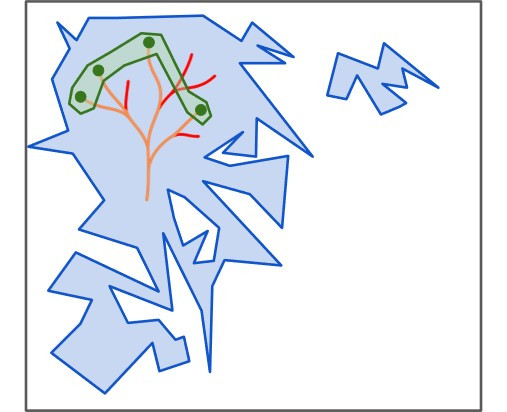
さまざまなタンパク質配列空間を抽象的に示した、縮尺が正確ではない視覚的表現。大きな箱は、すべてのアミノ酸配列の可能性(鎖の長さをLとした場合、約20L通りの組み合わせ)を表している。より小さな青い領域は、機能的なタンパク質を形成する配列を表しており、さらに小さな緑色の領域は、これまでに存在が確認されている配列を示している。金色の線はタンパク質の進化経路を表し、赤色の線は絶滅した進化経路(すなわち、現代の生物多様性には存在しないと考えられるもの)を表している。© イサコバほか
<関連情報>
- https://www.oist.jp/ja/news-center/news/2026/3/26/identifying-limits-protein-evolution
- https://www.pnas.org/doi/10.1073/pnas.2532018123
共通祖先からの派生は、タンパク質配列空間の探索を制限する Descent from a common ancestor restricts exploration of protein sequence space
Lada H. Isakova, Elizaveta Streltsova, Olga O. Bochkareva, +1 , and Fyodor A. Kondrashov
Proceedings of the National Academy of Sciences Published:March 31, 2026
DOI:https://doi.org/10.1073/pnas.2532018123
Significance
Are natural protein sequences representative of all possible sequences that are functional? The sequence space is immense but proteins have been evolving for billions of years, so much of the possible functional space may have already been explored. We find that because sequence evolution of homologous proteins starts from a single common ancestor, protein sequence diversity has been limited to an extreme degree and even 4 billion years of evolution was insufficient to explore the functional sequence space. Protein engineering models that learn only from natural protein sequences may be limited in their ability to predict sequences outside the explored sequence space and empirical approaches that explore the unnatural sequence space may be necessary to fully realize their potential.
Abstract
How functional protein sequences are distributed in sequence space is fundamentally important for evolutionary theory and protein design, particularly if a large diversity of protein functions are hidden in evolutionarily unexplored areas of the sequence space. However, this question is understudied in part because experimental and computational studies use extant sequences as a starting point to study sequence space. Here, we study whether extant sequences are representative of the entire functional sequence space. Across thousands of protein families from vertebrates and bacteria we calculate the dimensionality and the volume of sequence space occupied by extant homologs. We find that the observed dimensionality and volume of extant sequence space are minuscule, many orders of magnitude smaller than what we estimated using a model of protein evolution. Simulating sequence evolution we then quantify the impact of phylogeny, selection, and epistasis on restricting the evolutionary exploration of sequence space. We find that sequence evolution from a single common ancestor, or a single point of origin in sequence space, is by far the largest limiting factor that reduces the dimensionality and volume of extant sequence space. These results indicate that there are vast areas of functional sequence space that have not been explored in evolution because of the excessive restrictions on natural exploration of the protein sequence space imposed by the point of origin effect. We suggest that protein design methods that rely on extant sequences may be limited in their ability to discover truly novel functions.


