2026-05-08 熊本大学
<関連情報>
- https://www.kumamoto-u.ac.jp/whatsnew/seimei-sentankenkyu/20260508
- https://www.kumamoto-u.ac.jp/daigakujouhou/kouhou/pressrelease/wgt3jw/release260508.pdf
- https://academic.oup.com/nar/advance-article/doi/10.1093/nar/gkag378/8664501
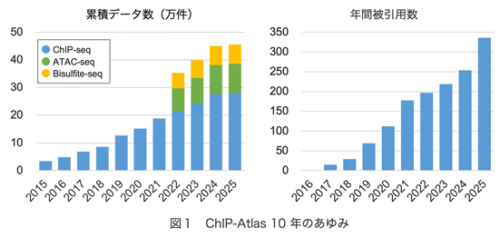
ChIP-Atlas 2025アップデート:エピゲノムランドスケープを探索するデータマイニングプラットフォームの10周年 ChIP-Atlas 2025 update: 10-year anniversary of a data-mining platform for exploring epigenomic landscape
Zhaonan Zou ,Tazro Ohta ,Takeya Kasukawa ,Shinya Oki
Nucleic Acids Research Published:29 April 2026
DOI:https://doi.org/10.1093/nar/gkag378
Abstract
ChIP-Atlas (https://chip-atlas.org/) is a data-mining platform that systematically processes and integrates public epigenomic profiling data, including ChIP-seq, ATAC-seq, DNase-seq, and Bisulfite-seq, together with curated sample metadata, currently encompassing >460 000 experiments across multiple model organisms. In this 2025 update, an experiment-level quality control framework was embedded in every experiment page, enabling intuitive assessment of data reliability and representativeness. In addition, a parametric analysis of gene set enrichment-based module was implemented, allowing enrichment analysis directly from continuous RNA-seq count tables without relying on threshold-defined, discrete gene lists, thereby helping elucidate underlying gene regulatory mechanisms. Combined with sustained data expansion, these updates advance ChIP-Atlas into a more quantitative and flexible platform for exploring the epigenomic landscape in multicellular organisms, marking its 10th anniversary.