2026-03-30 国立遺伝学研究所
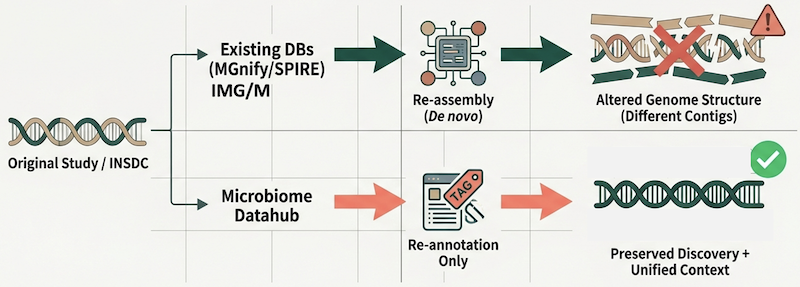
Microbiome Datahubと他の大規模MAGデータベースにおけるMAGの違い
<関連情報>
- https://www.nig.ac.jp/nig/ja/2026/03/research-highlights_ja/pr20260330.html
- https://www.nig.ac.jp/nig/images/research_highlights/PR20260330.pdf
- https://link.springer.com/article/10.1186/s40168-026-02385-x
マイクロバイオームデータハブ:環境メタデータ、分類、機能アノテーションを統合した、包括的なメタゲノムアセンブルゲノムデータセットのためのオープンアクセスプラットフォーム Microbiome Datahub: an open-access platform integrating environmental metadata, taxonomy, and functional annotation for comprehensive metagenome-assembled genome datasets
Hiroshi Mori,Takatomo Fujisawa,Koichi Higashi,Yasuhiro Tanizawa,Zenichi Nakagawa,Hiroyo Nishide,Masaki Fujiyoshi,Yasukazu Nakamura,Ikuo Uchiyama,Motomu Matsui & Takuji Yamada
Microbiome Published:16 March 2026
DOI:https://doi.org/10.1186/s40168-026-02385-x Unedited version
Abstract
Background
Metagenome-assembled genomes (MAGs) provide crucial insights into the genomic diversity of uncultured microbes. However, MAG datasets deposited in public repositories such as INSDC are often difficult to reuse due to heterogeneous quality, inconsistent taxonomic and functional annotations, and insufficiently curated environmental metadata. While secondary MAG databases such as MGnify, IMG/M, and SPIRE provide standardized resources, they reconstruct MAGs de novo from public metagenomic reads and therefore do not represent the original MAGs reported in publications.
Results
To address this gap, we developed Microbiome Datahub, an open-access platform that systematically aggregates and re-annotates original MAGs from INSDC. We collected 214,427 MAGs, predicted genes by DFAST, performed quality assessment with CheckM, standardized taxonomic assignments with GTDB-Tk, inferred 27 phenotypic traits using Bac2Feature, assigned proteins to MBGD ortholog clusters and KEGG Orthology IDs using PZLAST, and annotated environmental metadata with the Metagenome and Microbes Environmental Ontology. Across these MAGs, the average completeness was 80.5% and contamination 1.8%; notably, the most frequent values were >95% completeness and <1% contamination, indicating that the majority of MAGs are of high quality. Comparative analyses showed that Microbiome Datahub provides phylogenetically and environmentally diverse MAGs: while the majority originated from vertebrate gut environments, a substantial number were also recovered from other habitats such as groundwater, including nearly 10,000 MAGs from the Patescibacteria. Inference of 27 phenotypic traits, including optimum growth temperature, further revealed ecological differentiation across phyla. Protein clustering revealed 56 million identity 40% clusters, with the majority unique compared with MGnify and GlobDB, and ~19% of proteins unassigned to MBGD ortholog clusters, underscoring their novelty.
Conclusions
Microbiome Datahub integrates MAG genome sequences, gene and protein predictions, quality metrics, environmental and taxonomic annotations, ortholog cluster assignments, and phenotype predictions, all accessible via a web interface, API, and bulk downloads. By combining original MAGs with curated metadata and functional annotations, Microbiome Datahub constitutes a comprehensive and reusable resource that will accelerate microbiome and microbial genomics research.