2026-04-02 デラウェア大学(UD)
<関連情報>
- https://www.udel.edu/udaily/2026/april/synthetic-biology-microbes-biocontainment-aditya-kunjapur-amanda-forti/
- https://www.nature.com/articles/s41564-025-01999-5
- https://www.science.org/doi/10.1126/science.ado4537
非標準アミノ酸によって媒介される、人工的に設計された直交的かつ絶対的な細菌共生 Engineered orthogonal and obligate bacterial commensalism mediated by a non-standard amino acid
Amanda M. Forti,Michaela A. Jones,Defne N. Elbeyli,Neil D. Butler & Aditya M. Kunjapur
Nature Microbiology Published:01 May 2025
DOI:https://doi.org/10.1038/s41564-025-01999-5
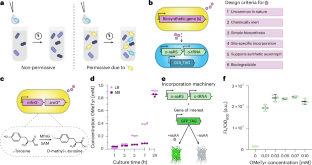
Abstract
Microorganisms can be genetically engineered for intrinsic biological containment based on synthetic chemical provision. However, reliance on an exogenous chemical limits the contexts where a contained microorganism could survive. Here we design an orthogonal obligate commensalism in Escherichia coli that autonomously creates environments permissive for survival of a partner microbe. We engineer one E. coli strain (the producer) to biosynthesize a non-standard amino acid (nsAA) from simple carbon sources through heterologous expression. We engineer a second E. coli strain (the utilizer) to rely on the same nsAA for growth as a synthetic auxotroph, with a 14-day escape rate of 2.8 × 10−9 escapees per colony-forming unit. Co-culture experiments show utilizer dependence on the producer, with no escape detected during co-inoculation of ~107 colony-forming units of utilizer and a non-producer E. coli strain. Dependence is maintained within a simplified synthetic maize root-associated community. This work provides ecological insights and presents a potential biocontainment strategy independent of an exogenous chemical.
抗原に化学的な目印を付ける 次世代生ワクチンは、硝化抗原の自律的な生産によって作られる Planting a chemical flag on antigens Next-generation live vaccines are created by autonomous production of nitrated antigens
Aditya M. Kunjapur
Science Published:4 Apr 2024
DOI:https://doi.org/10.1126/science.ado4537
Bacterial infections are a common cause of death across the globe and are an increasing threat as the prevalence of antibiotic resistance rises. Vaccines directed against bacterial pathogens prevent the spread of disease without the need for antibiotics and have an estimated global market size of 39.6 billion USD by 2030, with a compound annual growth rate of 7.7% (1). However, vaccines for bacterial pathogens are difficult to design owing to the ability of these organisms to hide their most potent antigens from the immune system (2). Weakened forms of live bacteria are some of the first vaccines developed and have some advantages as candidate vaccines compared to individual bacterial proteins (3). Yet, there are few options that balance safety and efficacy of live bacterial vaccines other than to control attenuation and dosage (4). My laboratory conducts several avenues of research to address this unmet need for improved bacterial vaccines as the antibiotic resistance crisis looms.