2026-03-18 カリフォルニア大学サンディエゴ校(UCSD)
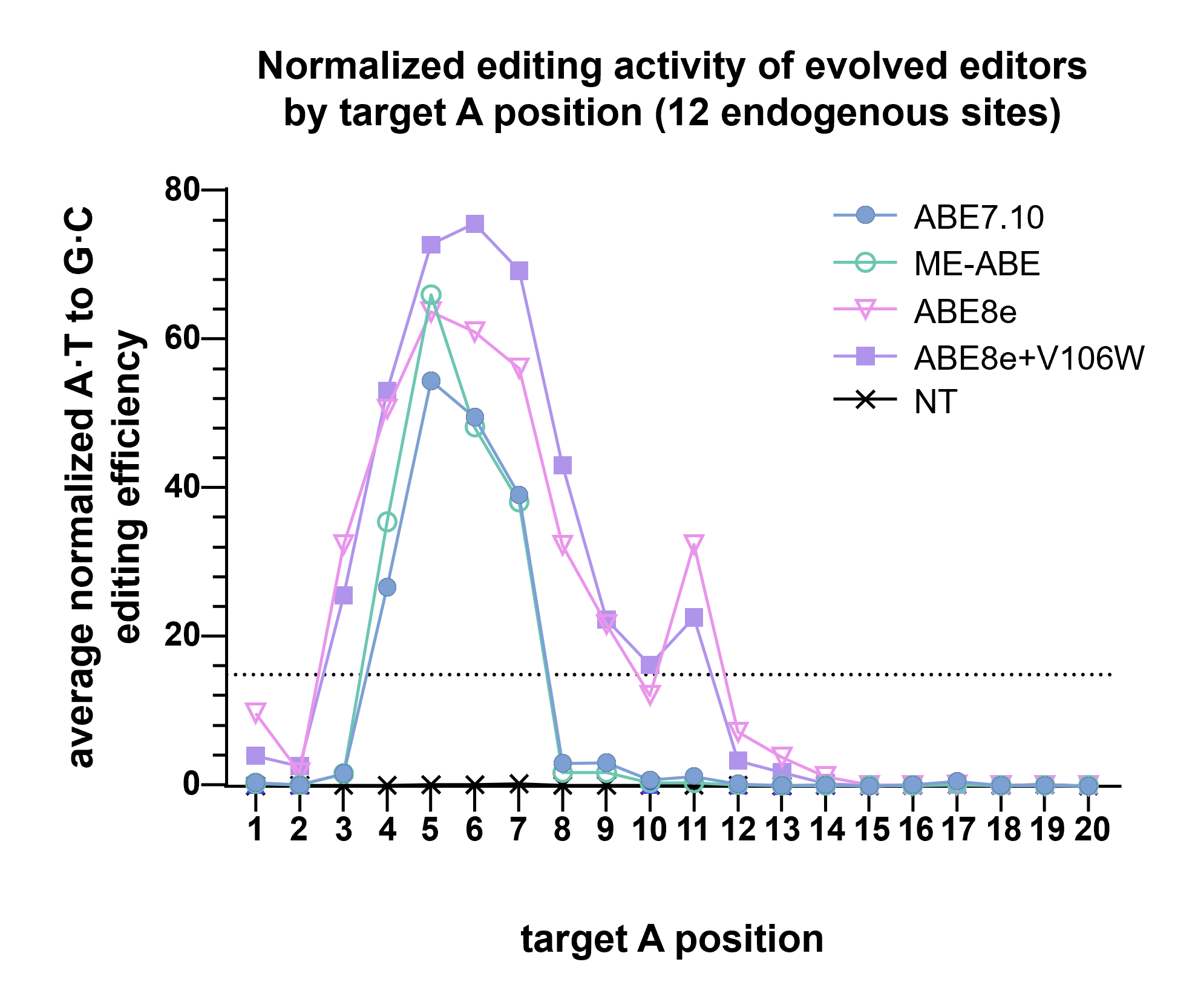
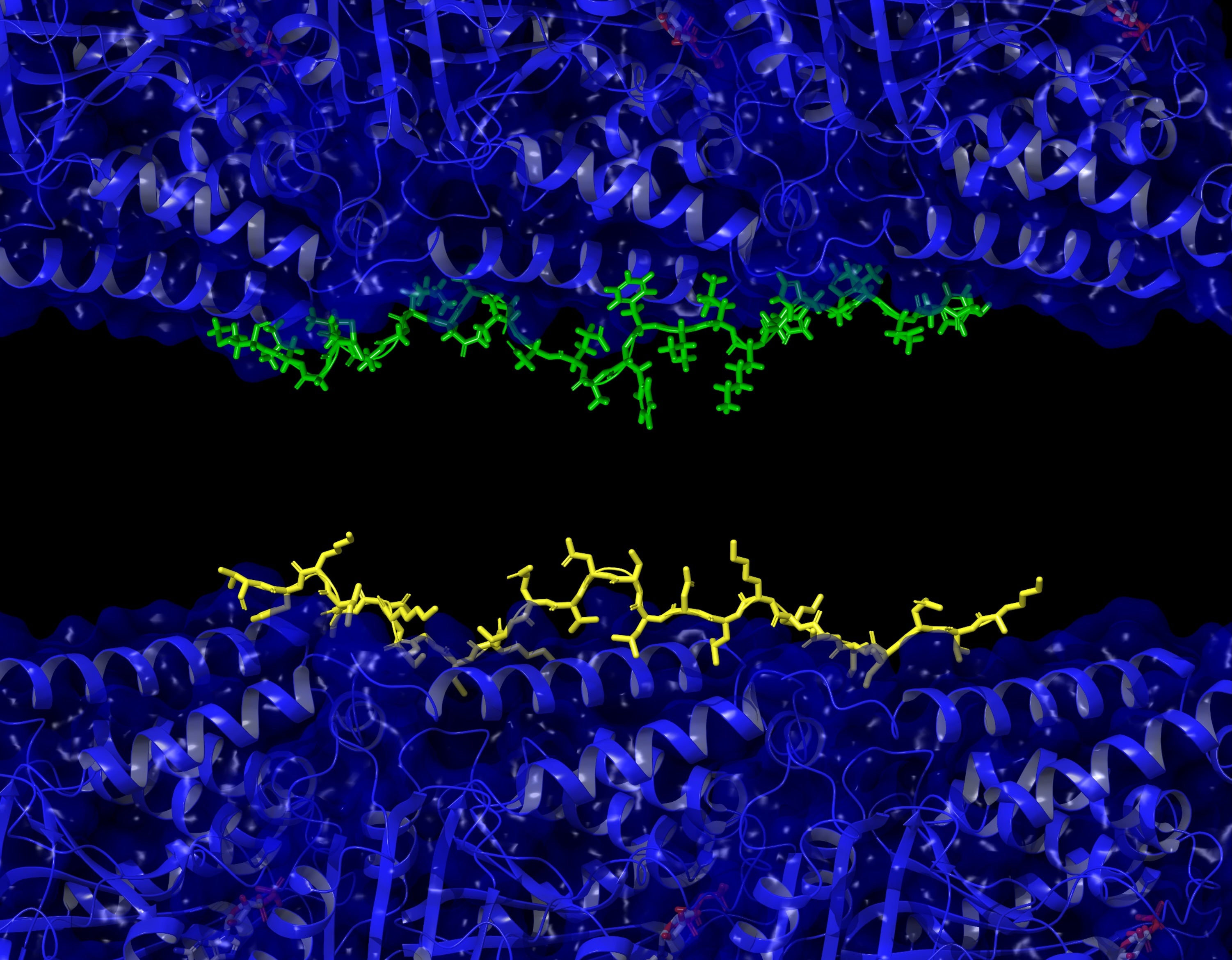
ABE 8 variants show a higher editing efficiency, but a larger editing window. ABE7.10 shows the opposite: a smaller window, but lower efficiency. ME-ABEs strike a balance, with small editing windows and high editing efficiency.
<関連情報>
- https://today.ucsd.edu/story/curing-the-bystander-effect
- https://www.nature.com/articles/s41587-026-03045-z
変異復帰解析によって生成された、精密かつ最小限の進化を遂げたアデニン塩基エディター Precise, minimally evolved adenine base editors generated through mutation reversion analysis
Mallory Evanoff,Sanjana Korpal,Zachary D. Krill,Quinn T. Cowan & Alexis C. Komor
Nature Biotechnology Published:18 March 2026
DOI:https://doi.org/10.1038/s41587-026-03045-z
Abstract
The initial development of adenine base editors (ABEs), which facilitate A•T to G•C base pair changes in the genome, used directed evolution to install 14 mutations into the wild-type deaminase TadA, producing the first-of-its-kind editor ABE7.10. Here we study the installed mutations’ impacts on TadA fitness using comprehensive reversion analysis and apply our results to engineer more efficient, precise editors. By measuring activity in both human and Escherichia coli host systems, we categorize mutations as critical, dispensable or host dependent. We show that up to five mutations can be reverted back to wild type, generating minimally evolved ABEs (ME-ABEs). ME-ABEs show narrow editing windows (similar to that of ABE7.10) and enhanced on-target editing (matching activities of the high-activity editor variants ABE8e and ABE8.20 in most sequence contexts) and exhibit low levels of guide-RNA-dependent and guide-RNA-independent off-target activity. ME-ABEs efficiently target six sites of clinical interest that had previously proved challenging to edit with ABE7.10, ABE8e or ABE8.20.