2026-03-30 マウントサイナイ医療システム (MSHS)
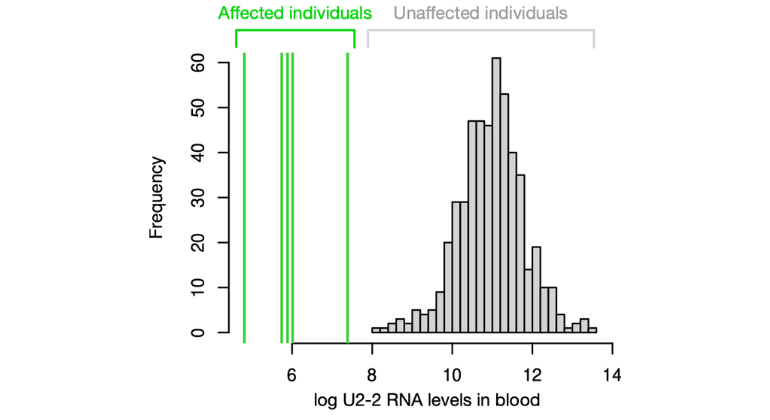
Histogram of log-scale U2-2 RNA levels in unaffected individuals (grey). The superimposed green lines denote five affected individuals with recessive ReNU2 syndrome. The stark difference between affected and unaffected individuals enables accurate diagnostics and points to gene replacement as a possible therapy. Adapted from Greene et al., Nature Genetics 2026.
<関連情報>
- https://www.mountsinai.org/about/newsroom/2026/researchers-identify-the-most-common-recessive-neurodevelopmental-disorder-ever-discovered
- https://www.nature.com/articles/s41588-026-02539-5
- https://www.nature.com/articles/s41588-025-02159-5
- https://www.nature.com/articles/s41591-024-03085-5
RNU2-2遺伝子の両アレル変異は、既知の最も一般的な劣性神経発達障害を引き起こす Biallelic variants in RNU2-2 cause the most prevalent known recessive neurodevelopmental disorder
Daniel Greene,Rodrigo Mendez,Jon Lees,Mafalda Barbosa,Alessandro Bruselles,Luigi Chiriatti,Federico Ferraro,Cecilia Mancini,Rachel Schot,Frank Sleutels,Enrico Bertini,Devon E. Bonner,Arjan Bouman,Alice S. Brooks,Thomas A. Cassini,Kimberly M. Ezell,Natalia Gomez-Ospina,Tjitske Kleefstra,Michael O’Donoghue,Lynette Rives,Vandana Shashi,Rebecca C. Spillmann,Mohamed Wafik,Undiagnosed Diseases Network,… Ernest Turro
Nature Published:30 March 2026
DOI:https://doi.org/10.1038/s41588-026-02539-5
Abstract
We recently showed that mutations in the snRNA genes RNU4-2 and RNU2-2 are prevalent causes of dominant neurodevelopmental disorders (NDDs). Here, by genetic association, we demonstrate the existence of a recessive form of RNU2-2 syndrome. We inferred a log Bayes factor for a recessive model of association of 18.2. Conditional on that model, 17 rare variants had a posterior probability of pathogenicity >0.8. This conservative threshold identified 18 probands and 5 affected siblings, each carrying two alleles in trans at these variants. A relaxed threshold of >0.6 identified a further 13 candidate probands. We identified nine further cases in replication collections. Affected individuals have intellectual disability, global developmental delay and seizures. Recessive RNU2-2 syndrome accounts for ~10% of families with a recessive NDD presently diagnosable by sequencing and affects ~60% as many families as the dominant RNU4-2-related NDD ReNU syndrome. The variants are predicted to destabilize stem loops and binding domains of U2-2 snRNA. Whole-blood RNA sequencing data showed a >90% reduction in the expression of pathogenic U2-2 alleles in biallelic cases and monoallelic carriers, albeit with wild-type compensation in carriers, pointing to a loss-of-expression mechanism.
小型核RNA遺伝子RNU2-2の変異は、顕著なてんかんを伴う重篤な神経発達障害を引き起こす Mutations in the small nuclear RNA gene RNU2-2 cause a severe neurodevelopmental disorder with prominent epilepsy
Daniel Greene,Koenraad De Wispelaere,Jon Lees,Marta Codina-Solà,Brynjar O. Jensson,Emma Hales,Andrea Katrinecz,Esther Nieto Molina,Sonia Pascoal,Rolph Pfundt,Rachel Schot,Marta Sevilla Porras,Frank Sleutels,Irene Valenzuela,Robin Wijngaard,Ignacio Arroyo Carrera,Giles Atton,Didac Casas-Alba,Deirdre Donnelly,Anna Duat Rodríguez,Bárbara Fernández Garoz,Nicola Foulds,Deyanira García-Navas Núñez,Elena González Alguacil,… Ernest Turro
Nature Genetics Published:10 April 2025
DOI:https://doi.org/10.1038/s41588-025-02159-5
Abstract
The major spliceosome includes five small nuclear RNA (snRNAs), U1, U2, U4, U5 and U6, each of which is encoded by multiple genes. We recently showed that mutations in RNU4-2, the gene that encodes the U4-2 snRNA, cause one of the most prevalent monogenic neurodevelopmental disorders. Here, we report that recurrent germline mutations in RNU2-2 (previously known as pseudogene RNU2-2P), a 191-bp gene that encodes the U2-2 snRNA, are responsible for a related disorder. By genetic association, we identified recurrent de novo single-nucleotide mutations at nucleotide positions 4 and 35 of RNU2-2 in nine cases. We replicated this finding in 16 additional cases, bringing the total to 25. We estimate that RNU2-2 syndrome has a prevalence of ~20% that of RNU4-2 syndrome. The disorder is characterized by intellectual disability, autistic behavior, microcephaly, hypotonia, epilepsy and hyperventilation. All cases display a severe and complex seizure phenotype. We found that U2-2 and canonical U2-1 were similarly expressed in blood. Despite mutant U2-2 being expressed in patient blood samples, we found no evidence of missplicing. Our findings cement the role of major spliceosomal snRNAs in the etiologies of neurodevelopmental disorders.
U4 snRNA遺伝子RNU4-2の変異は、最も一般的な単一遺伝子性神経発達障害の一つを引き起こす Mutations in the U4 snRNA gene RNU4-2 cause one of the most prevalent monogenic neurodevelopmental disorders
Daniel Greene,Chantal Thys,Ian R. Berry,Joanna Jarvis,Els Ortibus,Andrew D. Mumford,Kathleen Freson & Ernest Turro
Nature Medicine Published:31 May 2024
DOI:https://doi.org/10.1038/s41591-024-03085-5
Abstract
Most people with intellectual disability (ID) do not receive a molecular diagnosis following genetic testing. To identify new etiologies of ID, we performed a genetic association analysis comparing the burden of rare variants in 41,132 noncoding genes between 5,529 unrelated cases and 46,401 unrelated controls. RNU4-2, which encodes U4 small nuclear RNA, a critical component of the spliceosome, was the most strongly associated gene. We implicated de novo variants among 47 cases in two regions of RNU4-2 in the etiology of a syndrome characterized by ID, microcephaly, short stature, hypotonia, seizures and motor delay. We replicated this finding in three collections, bringing the number of unrelated cases to 73. Analysis of national genomic diagnostic data showed RNU4-2 to be a more common etiological gene for neurodevelopmental abnormality than any previously reported autosomal gene. Our findings add to the growing evidence of spliceosome dysfunction in the etiologies of neurological disorders.