2026-04-24 東京科学大学
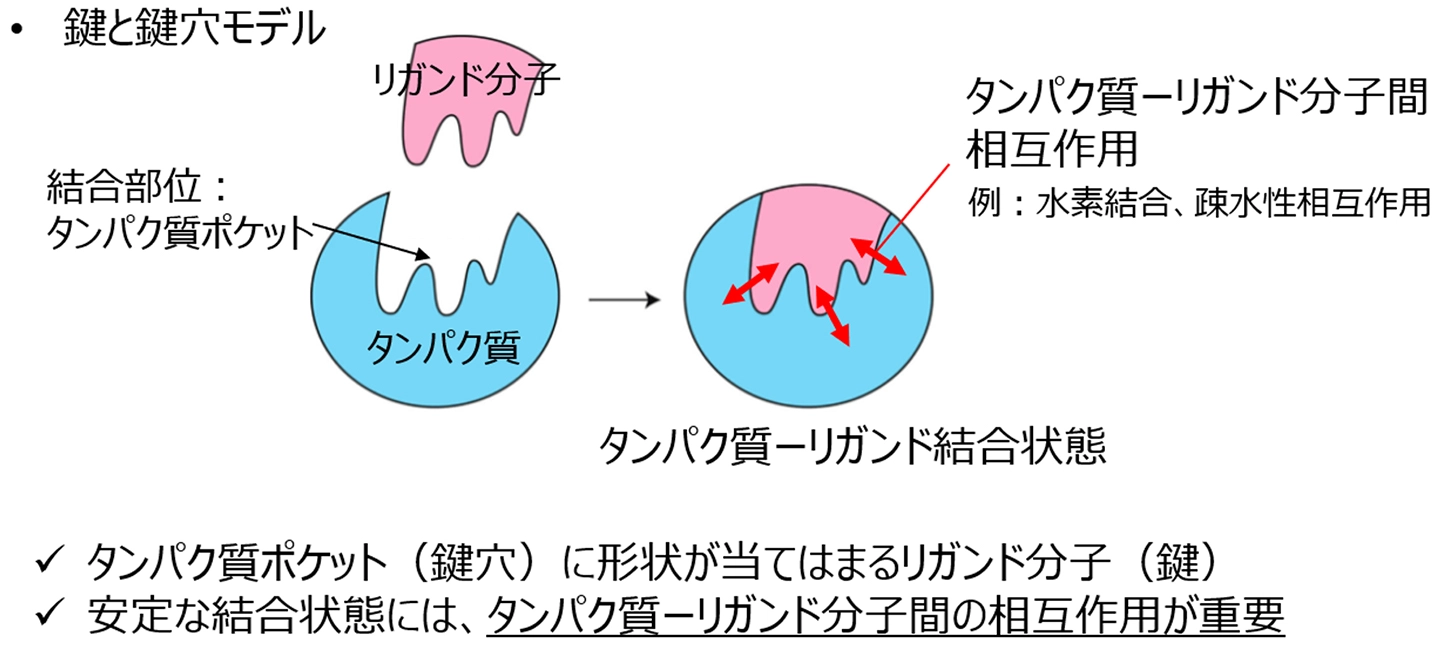
図1. 創薬研究における鍵と鍵穴モデル
<関連情報>
拡散モデルを用いた相互作用制約付き3D分子生成により、創薬のための構造ベースのファーマコフォアモデリングが可能になる Interaction-constrained 3D molecular generation using a diffusion model enables structure-based pharmacophore modeling for drug design
Masami Sako,Nobuaki Yasuo & Masakazu Sekijima
npj Drug Discovery Published:02 March 2026
DOI:https://doi.org/10.1038/s44386-026-00040-x
Abstract
A key challenge in structure-based drug design is generating three-dimensional molecules while preserving essential protein-ligand interactions. We propose DiffPharma, a structure-based pharmacophore modeling framework based on a conditional diffusion model, to generate molecules that satisfy specified interaction constraints. The proposed method incorporates a semantic fusion architecture that integrates multiple interaction-specific neural networks, each designed to capture distinct molecular interactions such as hydrogen bonds and hydrophobic interactions. Experimental results demonstrate that DiffPharma achieves a residue-level interaction similarity of up to 0.9, significantly outperforming baseline models. To assess the method’s generalizability, ligands were generated for AKT serine/threonine kinase 1 and serine β-lactamase, successfully preserving key interaction features. The effectiveness of the method is demonstrated through a practical case study targeting the SARS-CoV-2 main protease. Molecular dynamics simulations indicate that the generated molecules maintain both structural stability and key interactions comparable to those of a bioactive reference ligand. In addition, the molecular mechanics generalized Born surface area (MM/GBSA) calculations based on MD trajectories suggest that several generated molecules may exhibit relatively favorable binding tendencies compared with the reference. The implementation of the DiffPharma, including code and an execution environment on Google Colab, is available under the MIT license at GitHub: https://github.com/sekijima-lab/DiffPharma.