2026-03-03 ペンシルベニア州立大学(Penn State)
<関連情報>
- https://www.psu.edu/news/engineering/story/engineering-better-heartbeat
- https://pubs.aip.org/aip/cha/article-abstract/35/12/121105/3375085/Self-organizing-network-simulation-of-cardiac
- https://pubs.aip.org/aip/cha/article-abstract/35/4/041102/3343225/Self-organizing-network-simulation-of-cardiac
- https://pubs.aip.org/aip/cha/article-abstract/34/12/121102/3323315/Self-organizing-network-representation-of-human
心臓収縮ダイナミクスの自己組織化ネットワークシミュレーション Self-organizing network simulation of cardiac contraction dynamics
Runsang Liu;Hui Yang
Chaos Published:December 17 2025
DOI:https://doi.org/10.1063/5.0305451
Self-organizing network encodes and resembles structural geometry in the heart, offering a new pathway to study cardiac simulation. Hence, we have leveraged the sparsity of an adjacency matrix to design novel simulations of cardiac electrical dynamics on the self-organizing network. However, very little has been done to investigate network simulation of mechanical contraction dynamics. As a vertical step, this paper presents a new self-organizing network methodology for simulation modeling of cardiac contraction dynamics. To this end, we propose to model the self-organizing network as an interconnected spring–mass–damper system and further solve networked dynamic equations to simulate the orchestrated dynamics of mechanical contractions. The proposed methodology is evaluated and illustrated on both 2D cardiac tissues and the 3D heart. Experimental results show that the proposed methodology not only effectively models contraction dynamics in excitable media, but can also be flexibly extended to the whole heart.
心臓電気ダイナミクスの自己組織化ネットワークシミュレーション Self-organizing network simulation of cardiac electrical dynamics
Runsang Liu;Hui Yang
Chaos Published: April 10 2025
DOI:https://doi.org/10.1063/5.0261019
Network provides a low-dimensional representation of the heart through a sparse adjacency matrix, which ushers in a new opportunity to conduct cardiac simulation. We discovered that a self-organizing network encodes and resembles complex heart geometry. This, in turn, helps characterize the structure–function relationship of the heart through network theory. However, very little has been done to investigate the simulation of electrical activity on a self-organizing network. Thus, this paper presents a new self-organizing network approach for simulating cardiac electrical dynamics. We formulate and solve reaction–diffusion equations on the self-organizing network to simulate the propagation and turbulent behavior of electrical waves. The proposed methodology is evaluated and validated on both 2D cardiac tissues, consisting of healthy and infarcted cells, and the whole heart. Experimental results show that the proposed approach not only yields a compact network representation that resembles the heart geometry but also provides an effective simulation of spatiotemporal dynamics when benchmarking with traditional finite element method simulations.
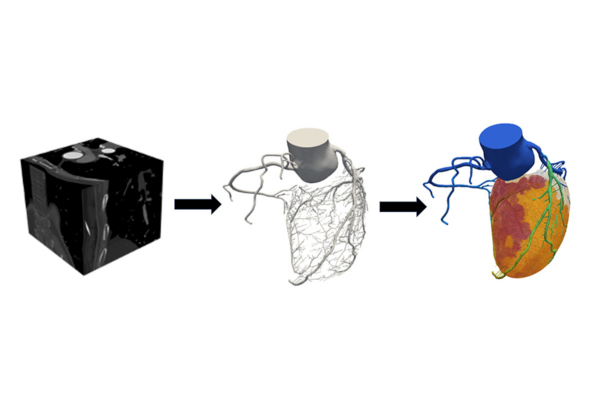
人間の心臓の自己組織化ネットワーク表現 Self-organizing network representation of human heart
Runsang Liu;Hui Yang
Chaos Published:December 02 2024
DOI:https://doi.org/10.1063/5.0243391
Network represents adjacent relationships, connections, and interactions among constituent elements in complex systems but often loses critical information about spatial configurations. However, structure–function relationships in biological systems, e.g., the human heart, are highly dependent on both connectivity relationships and geometric details. Therefore, this paper presents a new self-organizing approach to derive the geometric structure from a network representation of the heart. We propose to simulate the network as a physical system, where nodes are treated as charged particles and edges as springs and then let these nodes self-organize to reconstruct geometric details. Despite random initiations, this network evolves into a steady topology when its energy is minimized. This study addresses the open question, i.e., “whether a network representation can effectively resemble spatial geometry of a biological system,” thereby paving a stepstone to leverage network theory to investigate disease-altered biological functions.