2026-04-16 横浜市立大学
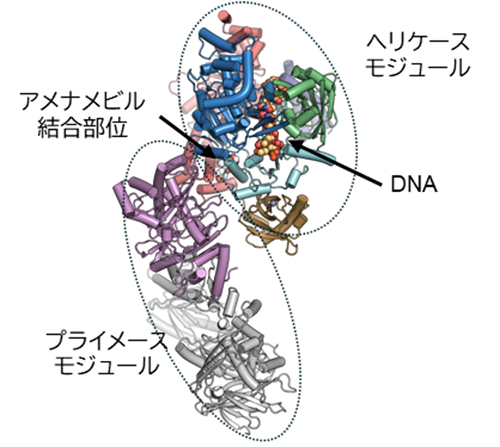
図1 単純ヘルペスウイルス由来HPCがDNAとアメナメビルを結合した構造。HPCは2つのモジュールに分けられ、DNAとアメナメビルは共にヘリケースモジュールに結合していた
<関連情報>
- https://www.yokohama-cu.ac.jp/res-portal/news/2026/20260416sengoku.html
- https://www.cell.com/cell-chemical-biology/abstract/S2451-9456(26)00076-0
ヒトヘルペスウイルス1型由来ヘリカーゼ-プライマーゼ複合体の阻害機構に関する構造的知見 Structural insights into inhibition mechanism of the helicase-primase complex from human herpesvirus 1
Ko Sato ∙ Hisashi Ishida ∙ Toma Miyagishi ∙ … ∙ Hidetoshi Kono ∙ Kaori Fukuzawa ∙ Toru Sengoku
Cell Chemical Biology Published:April 15, 2026
DOI:https://doi.org/10.1016/j.chembiol.2026.03.006
Significance
Herpesviruses are ubiquitous human pathogens responsible for a wide spectrum of diseases, including severe infections in immunocompromised individuals and associations with neurodegenerative and autoimmune disorders. Despite the clinical importance of these viruses, current antiviral therapies are largely limited to the α-subfamily, with few effective options for β- and γ-subfamilies. The helicase-primase complex (HPC) is a critical enzyme for viral DNA replication and a validated antiviral target, yet its structural basis and inhibition mechanisms have remained elusive, hindering the rational development of next-generation therapeutics. This study provides the first high-resolution cryo-EM structures of the human herpesvirus 1 (HHV1) HPC bound to single-stranded DNA and two clinically relevant inhibitors, amenamevir and pritelivir. These structures reveal the modular organization of the HPC and identify a shared allosteric pocket where both inhibitors bind. Structural, biochemical, and molecular dynamics analyses demonstrate that these inhibitors lock the helicase in an open, catalytically inactive conformation, thereby blocking ATP binding and halting DNA unwinding. Fragment molecular orbital (FMO) calculations further elucidate the molecular determinants of inhibitor binding, explaining the α-subfamily specificity and the distinct antiviral spectra of amenamevir and pritelivir. The integrated approach combining cryo-EM, molecular dynamics, and quantum chemical calculations not only clarifies the mechanism of action of current clinical inhibitors but also establishes a robust framework for the rational design of novel antivirals. By providing detailed insights into the structural and dynamic properties of the HPC and its inhibition, this work paves the way for the development of new drugs with tailored specificity and improved pharmacokinetic properties, potentially expanding therapeutic options to currently untreatable herpesvirus infections. These findings have broad implications for antiviral drug discovery and for understanding the molecular mechanisms underlying DNA replication in herpesviruses.
Highlights
- HHV1 helicase-primase complex has a defined architecture revealed by cryo-EM
- Amenamevir and pritelivir bind a common allosteric pocket in the complex
- Inhibitors lock helicase in an open, inactive state, blocking ATP binding
- α-subfamily selectivity and antiviral differences arise from inhibitor interactions
Summary
Human herpesviruses (HHVs) are widespread pathogens causing severe diseases, especially in immunocompromised individuals, and are linked to neurodegenerative and autoimmune disorders. Current antiviral therapies primarily target the α-subfamily, with limited options for β- and γ-subfamilies. The helicase-primase complex (HPC) is essential for viral DNA replication and represents a key antiviral target, but its structure and inhibition mechanisms were previously unclear. Here, cryo-EM structures of the HHV1 (α-subfamily) HPC bound to single-stranded DNA and two clinical inhibitors, amenamevir and pritelivir, are reported. The HPC features flexible helicase and primase modules, with both inhibitors binding a shared allosteric pocket in the helicase module. Structural, biochemical, and molecular dynamics analyses indicate that these inhibitors lock the helicase in an open, inactive conformation. Combined molecular dynamics and fragment molecular orbital analyses explain the α-subfamily selectivity and distinct antiviral spectra of these drugs. These findings provide a structural framework to guide the development of novel inhibitors targeting β- and γ-subfamily herpesviruses.


