2026-03-02 東大病院
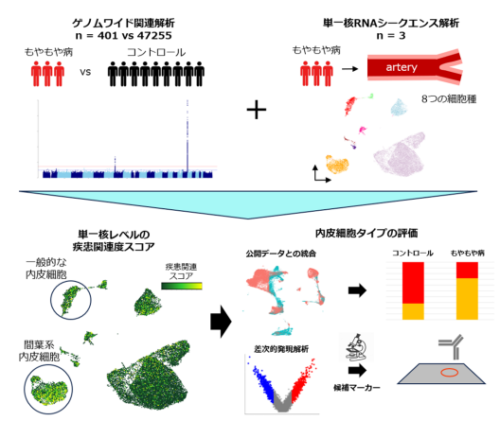
研究の全体像
<関連情報>
- https://www.h.u-tokyo.ac.jp/press/20260302.html
- https://www.h.u-tokyo.ac.jp/press/__icsFiles/afieldfile/2026/03/02/release_20260302.pdf
- https://www.ahajournals.org/doi/10.1161/STROKEAHA.125.053747
統合GWASとsnRNA-seqにより、もやもや病における間葉系血管内皮細胞のシグネチャーが明らかに Integrative GWAS and snRNA-seq Reveal a Mesenchymal-Like Endothelial Signature in Moyamoya Disease
Yudai Hirano, MD; Satoru Miyawaki, MD, PhD; Kyuto Sonehara, MD, PhD; Shinichi Namba, MD, PhD; Hirotaka Inoue, MD; Yuya Shirai, MD, PhD; Hideaki Imai, MD, PhD; …;on behalf of the BioBank Japan Project
Stroke Published 2 March 2026
DOI:https://doi.org/10.1161/STROKEAHA.125.053747
Abstract
BACKGROUND:
Moyamoya disease (MMD) has a strong genetic basis, with the rare RNF213 variant (rs112735431) representing a major risk factor, while the broader genetic architecture and disease-relevant vascular cell types remain incompletely understood.
METHODS:
We conducted a genome-wide association study in Japanese individuals (n=47 656; 401 MMD cases and 47 255 controls). Population-level features at MMD risk loci were examined by regional allele frequency and haplotype analyses. We performed single-nucleus RNA-seq of superficial temporal arteries from patients with MMD (n=3). Cell type–specific enrichment of genome-wide association study signals was assessed using the Single-Cell Disease Relevance Score. Endothelial signatures were validated by integration with publicly available single-cell data sets from controls (n=5) and immunohistochemistry for candidate markers (n=1).
RESULTS:
Beyond rs112735431, we identified a genome-wide significant signal in the HDAC9-TWIST1 region (P=3.3×10−14; odds ratio, 1.77). Conditional analysis on rs112735431 revealed a protective RNF213 missense variant, p.Asn1331Gly (rs8074015; P=3.7×10−9; odds ratio, 0.53), whose minor allele was mutually exclusive with rs112735431-A on haplotypes. Population analysis revealed geographic variation and extended haplotype structure of the rs112735431-A allele in Japan. Single-nucleus RNA-seq identified a mesenchymal-like endothelial cell (MEC) population with selective FN1 expression. Genome-wide association study–prioritized disease genes were strongly enriched in MECs. MECs showed mesenchymal pathway activation with a regulatory program distinct from canonical endothelial states. The proportion of MECs was markedly increased in MMD (72% versus 28% in controls), and FN1 expression in endothelial regions was confirmed by immunohistochemistry.
CONCLUSIONS:
Our findings identify a protective RNF213 variant that is mutually exclusive with the known rs112735431-A allele. Genetic risk converges on an MEC state markedly expanded in MMD.