2026-03-04 東京科学大学
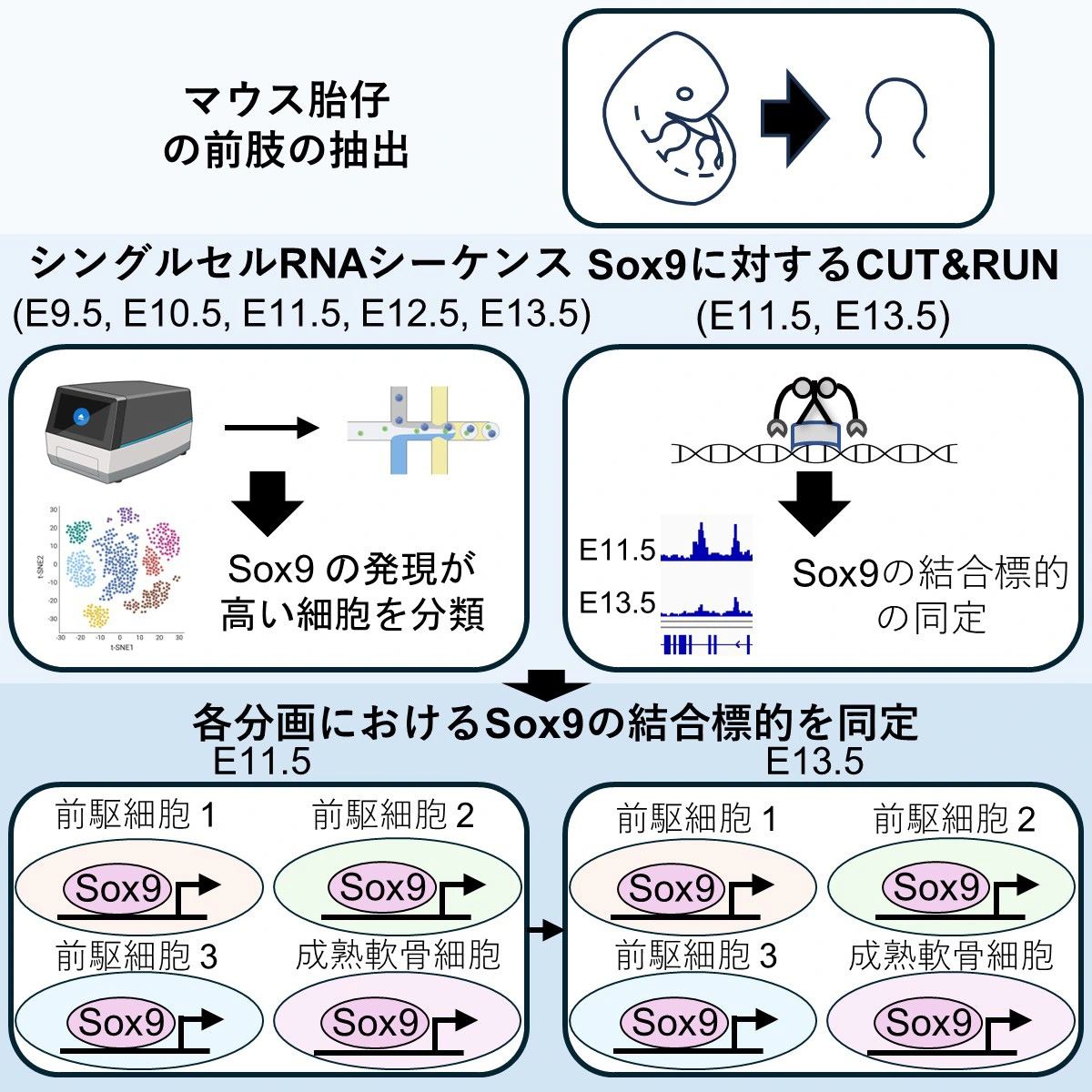
図1. 肢芽におけるSox9の時相・細胞特異的な標的の同定
<関連情報>
- https://www.isct.ac.jp/ja/news/w4ps6bktw06f#top
- https://www.cell.com/iscience/fulltext/S2589-0042(26)00449-9
マウス胎児肢芽におけるSox9による軟骨形成遺伝子プログラムの段階およびクラスター特異的制御 Stage- and cluster-specific regulation of chondrogenic gene programs by Sox9 in mouse embryonic limb buds
Masayasu Sega ∙ Yutaro Uchida ∙ Tomoki Chiba ∙ … ∙ Kohei Kita ∙ Naoyuki Miyasaka ∙ Hiroshi Asahara
iScience Published:February 18, 2026
DOI:https://doi.org/10.1016/j.isci.2026.115074
Highlights
- Three chondroprogenitor clusters and mature chondrocytes show stage-dependent dynamics
- CUT&RUN of Ty1-tagged Sox9 reveals stage- and cluster-specific chromatin occupancy
- Integration of CUT&RUN and scRNAseq identifies Sox9 targets in chondrocyte maturation
- Sox9 regulates transcriptional networks across cartilage developmental stages
Summary
Cartilage formation in the limb is initiated and sustained by Sox9, though how its regulatory outputs evolve across developmental time and cell states remains unclear. Here, we integrate single-cell RNA sequencing of mouse forelimb buds across five stages (E9.5–E13.5) with CUT&RUN profiling of Ty1-tagged Sox9 at E11.5 and E13.5. We identify four Sox9-high populations—three chondroprogenitor clusters and mature chondrocytes—with distinct dynamics, and RNA-velocity infers independent trajectories from each progenitor cluster to maturation. Sox9 chromatin occupancy shows a conserved motif signature but undergoes stage- and cluster-dependent reconfiguration, aligning with shifts in extracellular matrix, cell-cycle, and patterning programs. Integrative analysis links these binding differences with coordinated transcriptional changes, suggesting that Sox9 operates through context-associated regulatory modes rather than a single uniform program. Our study provides a stage-resolved regulatory map of Sox9-associated regulation during chondrogenesis and a resource for limb mesenchyme.