2026-03-18 スタンフォード大学
<関連情報>
- https://news.stanford.edu/stories/2026/03/protein-sequencing-method-dna-research
- https://www.nature.com/articles/s41587-026-03061-z
ペプチドをDNAに逆翻訳することによる単一分子ペプチド配列決定 Single-molecule peptide sequencing through reverse translation of peptides into DNA
Liwei Zheng,Yujia Sun,Linus A. Hein,Michael Eisenstein & Hyongsok Tom Soh
Nature Biotechnology Published:18 March 2026
DOI:https://doi.org/10.1038/s41587-026-03061-z
Abstract
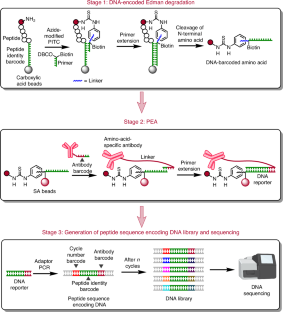
Despite advances in mass spectrometry and emerging single-molecule approaches, sequencing peptides at the single-molecule level remains a central challenge in proteomics. Here we present a ‘reverse translation’ strategy that enables single-molecule peptide sequencing with single-amino-acid resolution. In this approach, peptides undergo a modified Edman degradation that iteratively releases N-terminal amino acids tagged with peptide-specific DNA barcodes. Antibody-mediated proximity extension assays identify these barcoded amino acids and generate PCR-amplifiable DNA reporters that record the identity, position and originating peptide of each amino acid. The resulting DNA library is directly read by high-throughput sequencing, converting peptide sequences into digital DNA outputs. Using this approach, we demonstrate true single-molecule peptide sequencing, achieving full sequence coverage in millions of reads and accurate differentiation of both native and post-translationally modified peptides. These results establish a framework that redefines protein sequencing as a DNA sequencing problem and lays the foundation for high-throughput, de novo single-molecule protein sequencing.

