2026-04-13 米国国立衛生研究所(NIH)
<関連情報>
- https://www.nih.gov/news-events/news-releases/nih-funded-breakthrough-shrinks-crispr-precision-delivery-body
- https://www.nature.com/articles/s41594-026-01788-6
Cas12fオルソログの比較解析により、ゲノム編集効率向上を支えるメカニズム的特徴が明らかになった Comparative characterization of Cas12f orthologs reveals mechanistic features underlying enhanced genome editing efficiency
Kaoling Guan,Rodrigo Fregoso Ocampo,Paula B. Matheus Carnevali,Cindy J. Castelle,Liliana Gonzalez-Osorio,Dominic T. Castanzo,Nicole C. Thomas,Molly Brothers,Tyler L. Dangerfield,Matthew M. Hooper,Madeline S. West,Nathan M. Appleby,Isabella Krudop,Rebecca C. Lamothe,Daniela S. Aliaga Goltsman,Lisa M. Alexander,Cristina N. Butterfield,Kenneth A. Johnson,Christopher T. Brown & David W. Taylor
Nature Structural & Molecular Biology Published:13 April 2026
DOI:https://doi.org/10.1038/s41594-026-01788-6
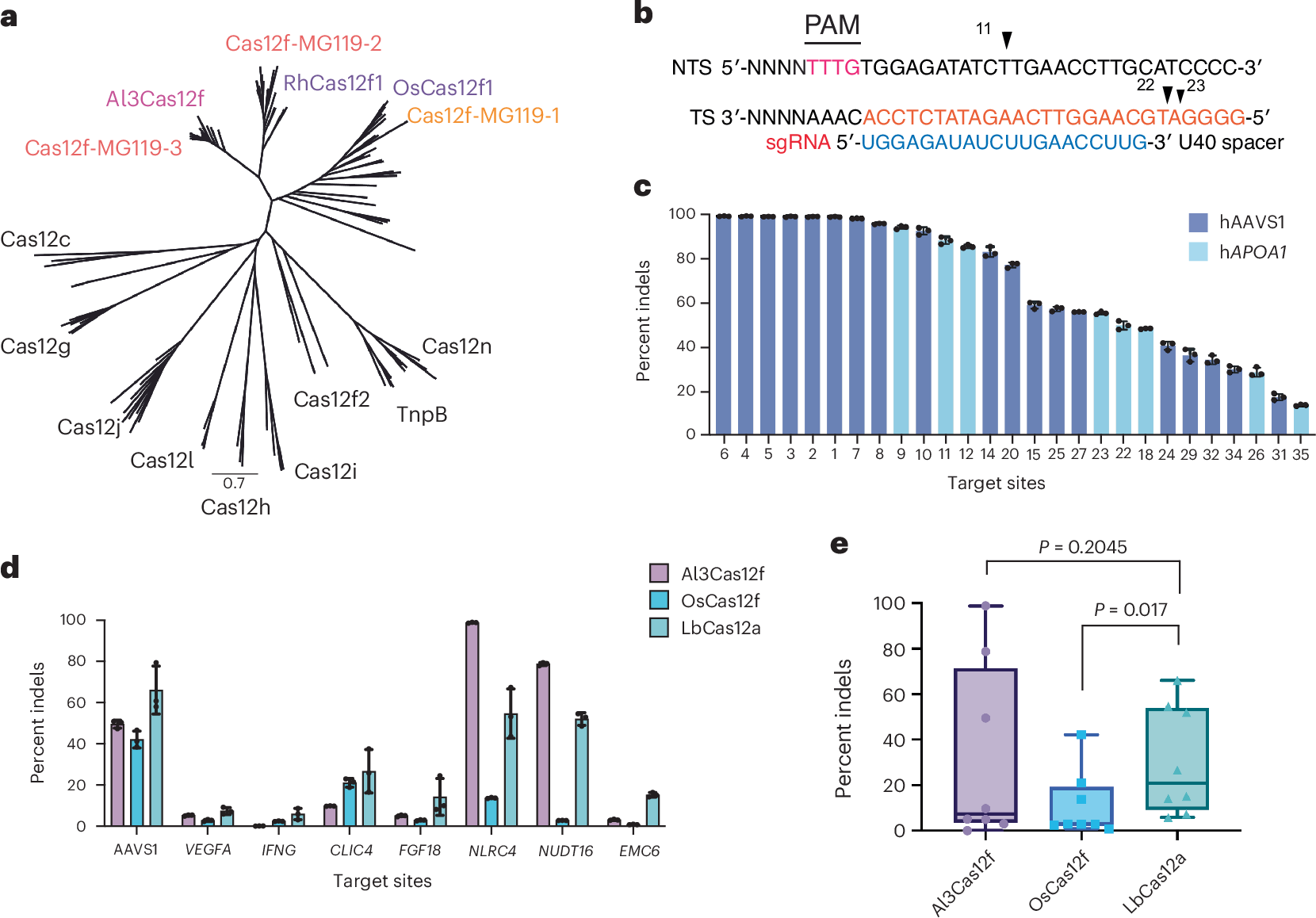
Abstract
Miniature CRISPR–Cas12f nucleases are attractive candidates for therapeutic genome editing because of their compact size and compatibility with adeno-associated virus (AAV) delivery. However, editing efficiencies in mammalian cells are lower than those of larger systems. The extensive phylogenetic diversity of Cas12f suggests unexplored mechanistic variation with the potential for optimization. Here we identify and characterize a naturally occurring Cas12f ortholog discovered through metagenomics, Alistipes sp. Cas12f (Al3Cas12f), which supports robust genome editing in human cells. Through structural, biochemical and kinetic analyses, we compare Al3Cas12f to two recently described orthologs, Oscillibacter sp. Cas12f and Ruminiclostridium herbifermentans Cas12f. These orthologs present divergent architectures and regulatory features governing protospacer-adjacent motif recognition, guide RNA (gRNA) binding, dimerization and DNA cleavage. Notably, Al3Cas12f achieves efficient R-loop formation through a stable dimer interface and a naturally optimized gRNA. Leveraging these structural insights, we generate an engineered Al3Cas12f variant (RKK) that increases editing and improves activity across several tested genomic loci. By overcoming locus-dependent variability and an apparent potency threshold, this engineered compact editor seems to expand the feasibility of low-dose, AAV-compatible therapeutic genome editing. Our results elucidate mechanistic determinants of Cas12f activity and offer a framework for engineering compact genome editors that may bear therapeutic potential.