2026-03-13 東京科学大学
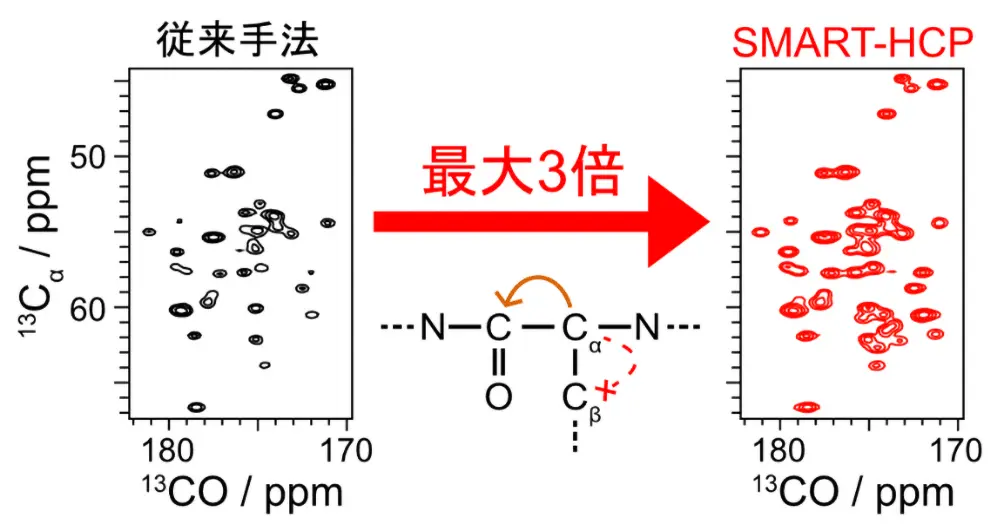
図1. プロテインG B1ドメインの13Cα–13CO2次元NMRスペクトル。左は従来手法であるDREAM法、右は本研究で開発したSMART-HCP法を使い、13Cα–13CO間で磁化移動を行った。SMART-HCP法で測定した信号感度は最大で3倍(平均1.7倍)大きくなった。従来手法では、信号感度を3倍にするために32≃9倍の実験時間が必要となる。
<関連情報>
- https://www.isct.ac.jp/ja/news/yuxjkk0364a8
- https://pubs.acs.org/doi/10.1021/acsmeasuresciau.5c00103
超高速MASを用いた高次元タンパク質固体NMRにおける13C – 13C偏極移動障壁の打破 Breaking the 13C–13C Polarization Transfer Barrier for High-Dimensional Protein Solid-State NMR with Ultra-Fast MAS
Tatsuya Matsunaga,Tsukito So,Ryo Takahashi,Yoshiki Shigemitsu,and Yoshitaka Ishii
ACS Measurement Science Au Published: February 2, 2026
DOI:https://doi.org/10.1021/acsmeasuresciau.5c00103
Abstract
Recent advances in 1H-detected solid-state NMR spectroscopy (SSNMR) using ultra-fast magic angle spinning (UFMAS) at frequencies above 60 kHz potentially facilitate high-dimensional SSNMR (HD-SSNMR) for protein analysis. A major limitation of HD-SSNMR is the exponential signal loss that occurs during successive polarization transfers. To overcome this bottleneck in HD-SSNMR for the most problematic 13C–13C transfers, we introduce a simple yet exceptionally efficient homonuclear cross-polarization (HCP) scheme called SeMi-selective Adiabatic Recoupling Transfer with HCP (SMART-HCP). We demonstrate that SMART-HCP with UFMAS at 90 kHz achieved nearly complete transfer with an efficiency of 76% from 13CO to 13Cα and 70% from 13Cα to 13CO for uniformly 13C,15N-labeled l-alanine. Semiselective HCP was achieved by optimizing radio frequency (RF)-offset frequency and amplitude modulations via a simple graphical method. For uniformly 13C,15N-labeled immunoglobulin-binding protein G (GB1) proteins, compared with the conventional dipolar recoupling enhancement through amplitude modulation (DREAM) scheme, SMART-HCP enhanced 13CO–13Cα transfers up to ∼3-fold (average 1.7-fold) for nonglycine residues, thereby accelerating various protein SSNMR experiments, including HD-SSNMR, by up to ∼9-fold. Our 3D 1H-detected (H)CACO(N)H SSNMR spectra of the GB1 sample suggest that with the SMART-HCP method a usually time-consuming 3D protein SSNMR experiment can be achieved within 3.5 h for a trace amount of the protein sample (∼200 μg). Besides biological applications, this method is likely applicable to 13C SSNMR analysis of a wide range of samples, such as polymers, peptide-based pharmacological agents, and other solid organic materials.