2026-04-29 ミュンヘン大学(LMU)
<関連情報>
- https://www.lmu.de/en/newsroom/news-overview/news/how-the-embryonal-epigenome-organizes-itself-01c3e9f3.html
- https://www.nature.com/articles/s41567-026-03263-x
胚性エピゲノム形成におけるスケーリングと自己相似性 Scaling and self-similarity in the formation of the embryonic epigenome
Fabrizio Olmeda,Tim Lohoff,Ioannis Kafetzopoulos,Stephen J. Clark,Laura Benson,Fatima Santos,Felix Krueger,Simon Walker,Wolf Reik & Steffen Rulands
Nature Physics Published:29 April 2026
DOI:https://doi.org/10.1038/s41567-026-03263-x
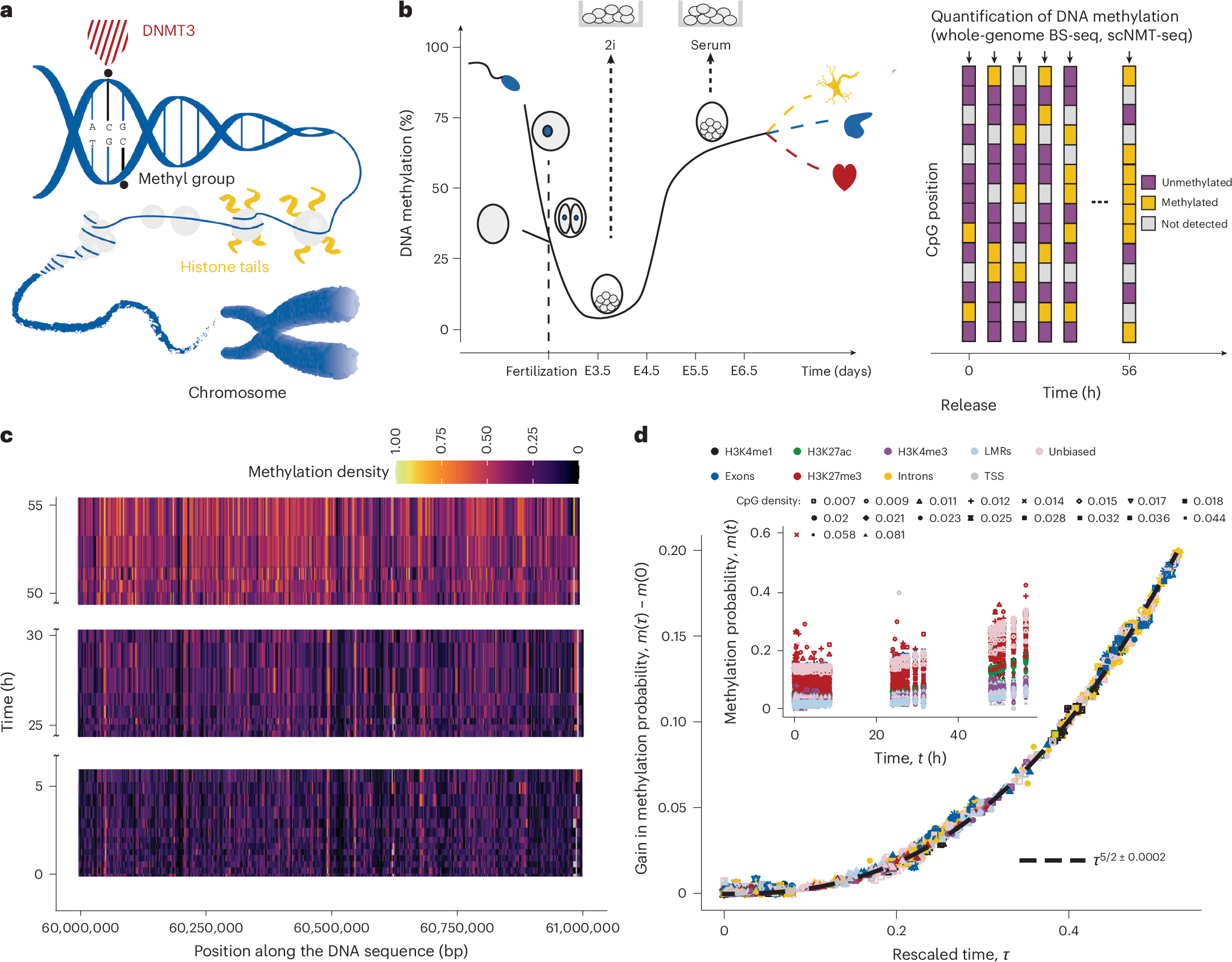
Abstract
The development of complex tissues relies on the precise assignment of cell identity. At the molecular scale, this process depends on the deposition of epigenetic modifications—such as methylation—that are regulated by complex biochemical networks and occur at specific regions on the DNA and chromatin. Here we show that despite the complexity of epigenetic regulation, dynamical scaling and self-similarity of DNA methylation marks emerge in embryonic development. Drawing on single-cell multi-omics experiments, super-resolution microscopy and statistical physics, we demonstrate that these phenomena originate in dynamical feedback between DNA methylation and the formation of nanoscale dynamic chromatin aggregates. These nanoscale processes lead to genome-wide increase in DNA methylation marks following a power law and self-similar correlation functions. Using this framework, we identify methylation patterns that precede gene expression changes in embryonic symmetry breaking. Our work identifies linear sequencing measurements as a laboratory to study mesoscopic biophysical processes in vivo.